Gut microbiota dysbiosis orchestrates vitiligo-related oxidative stress through the metabolite hippuric acid
- PMID: 40329424
- PMCID: PMC12054231
- DOI: 10.1186/s40168-025-02102-0
Gut microbiota dysbiosis orchestrates vitiligo-related oxidative stress through the metabolite hippuric acid
Abstract
Background: Vitiligo, a depigmenting autoimmune skin disease characterized by melanocyte dysfunction or death, is known to be associated with an imbalance in gut microbiota. Oxidative stress plays a critical role in the pathogenesis of vitiligo. However, the complex promising interaction between abnormal accumulation of reactive oxygen species (ROS) in the skin and gut microbiota has remained unclear.
Results: Here, we compared transcriptome data of vitiligo lesions and normal skin and identified a high expression of oxidative stress-related genes in vitiligo lesions. We also established a vitiligo mouse model and found that the presence of gut microbiota influenced the expression of ROS-related genes. Depletion of gut microbiota reduced abnormal ROS accumulation and mitochondrial abnormalities in melanocytes, significantly improving depigmentation. Our findings from manipulating gut microbiota through cohousing, fecal microbiota transplantation (FMT), and probiotic supplementation showed that transferring gut microbiota from mice with severe vitiligo-like phenotypes exacerbated skin depigmentation while probiotics inhibited its progression. Targeted metabolomics of fecal, serum, and skin tissues revealed gut microbiota-dependent accumulation of hippuric acid, mediating excessive ROS in the skin. Elevated serum hippuric acid levels were also confirmed in vitiligo patients. Additionally, a microbiota-dependent increase in intestinal permeability in vitiligo mice mediated elevated hippuric acid levels, and we found that hippuric acid could directly bind to ROS-related proteins (NOS2 and MAPK14).
Conclusions: Our results suggested the important role of gut microbiota in regulating vitiligo phenotypes and oxidative stress. We identified hippuric acid, a gut microbiota-host co-metabolite, as a critical mediator of oxidative stress in vitiligo skin and its binding targets (NOS2 and MAPK14), resulting in oxidative stress. Validation in a small human cohort suggested that hippuric acid could serve as a novel diagnostic biomarker and therapeutic target for vitiligo. These findings provided new insights into how gut microbiota regulates skin oxidative stress in vitiligo and suggested potential treatment strategies for the disease. Video Abstract.
Keywords: Gut microbiota; Hippuric acid; Oxidative stress; Vitiligo.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The experimental animals used in this study were adult male mice with a pure C57BL/6 J background, acquired from the Experimental Animal Center of the Fourth Military Medical University. All procedures were approved by the Fourth Military Medical University and adhered to all relevant ethical regulations. The mice were housed under a 12-h light/dark cycle at 22–25 °C with free access to water and food in environmentally controlled conditions. The UPLC-MS/MS analysis and ELISA analysis of serum from both advanced vitiligo patients and healthy volunteers were reviewed and approved by the Ethics Committee of the Air Force Medical Center (No. 2022–188-PJ01). All participants provided written informed consent for sample collection and data analysis. Prior to giving consent, participants were informed about the goals and related experimental procedures of the study. Consent for publication: All authors have approved the publication of this manuscript. Competing interests: The authors declare no competing interests.
Figures






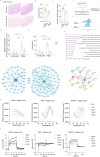
References
-
- Picardo M, Dell’Anna ML, Ezzedine K, Hamzavi I, Harris JE, Parsad D, Taieb A. Vitiligo. Nat Rev Dis Primers. 2015;1:15011. - PubMed
-
- Ezzedine K, Eleftheriadou V, Whitton M, van Geel N. Vitiligo. Lancet. 2015;386(9988):74–84. - PubMed
-
- Frisoli ML, Essien K, Harris JE. Vitiligo: mechanisms of pathogenesis and treatment. Annu Rev Immunol. 2020;38:621–48. - PubMed
-
- Rodrigues M, Ezzedine K, Hamzavi I, Pandya AG, Harris JE. Current and emerging treatments for vitiligo. J Am Acad Dermatol. 2017;77(1):17–29. - PubMed
-
- Kovacs SO. Vitiligo. J Am Acad Dermatol. 1998;38(5 Pt 1):647–66; quiz 667–648. - PubMed
MeSH terms
Substances
Grants and funding
- 82203909/National Natural Science Foundation of China grant
- 82100513/National Natural Science Foundation of China grant
- GZC20233592/Postdoctoral Fellowship Program of CPSF
- 22BJQN003/Youth Talent Support Program of Air Force Medical Center
- 21ZT10/Boost Program for Young Doctor of Air Force Medical Center
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

