The First Complete Chloroplast Genome of Spider Flower (Cleome houtteana) Providing a Genetic Resource for Understanding Cleomaceae Evolution
- PMID: 40332020
- PMCID: PMC12027348
- DOI: 10.3390/ijms26083527
The First Complete Chloroplast Genome of Spider Flower (Cleome houtteana) Providing a Genetic Resource for Understanding Cleomaceae Evolution
Abstract
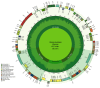
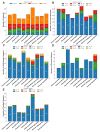
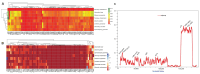
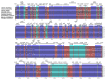
In the present study, the sequencing and analysis of the complete chloroplast genome of Cleome houtteana and its comparison with related species in the Cleomaceae family were carried out. The genome spans 157,714 base pairs (bp) and follows the typical chloroplast structure, consisting of a large single-copy (LSC) region (87,506 bp), a small single-copy (SSC) region (18,598 bp), and two inverted repeats (IRs) (25,805 bp each). We identified a total of 129 genes, including 84 protein-coding genes, 8 ribosomal RNA (rRNA) genes, and 37 transfer RNA (tRNA) genes. Our analysis of simple sequence repeats (SSRs) and repetitive elements revealed 91 SSRs, with a high number of A/T-rich mononucleotide repeats, which are common in chloroplast genomes. We also observed forward, palindromic, and tandem repeats, which are known to play roles in genome stability and evolution. When comparing C. houtteana with its relatives, we identified several highly variable regions, including ycf1, ycf2, and trnH-psbA, marking them as propitious molecular markers for the identification of species as well as phylogenetic studies. We examined the inverted repeat (IR) boundaries and found minor shifts in comparison to the other species, particularly in the ycf1 gene region, which is a known hotspot for evolutionary changes. Additionally, our analysis of selective pressures (Ka/Ks ratios) showed that most genes are under strong purifying selection, preserving their essential functions. A sliding window analysis of nucleotide diversity (Pi) identified several regions with high variability, such as trnH-psbA, ycf1, ndhI-ndhG, and trnL-ndhF, highlighting their potential for use in evolutionary and population studies. Finally, our phylogenetic analysis, using complete chloroplast genomes from species within Cleomaceae, Brassicaceae, and Capparaceae, confirmed that C. houtteana belongs within the Cleomaceae family. It showed a close evolutionary relationship with Tarenaya hassleriana and Sieruela rutidosperma, supporting previous taxonomic classifications. The findings from the current research offer invaluable insights regarding genomic structure, evolutionary adaptations, and phylogenetic relationships of C. houtteana, providing a foundation for future research on species evolution, taxonomy, and conservation within the Cleomaceae family.
Keywords: SSRs; Ycf1 gene; chloroplast genome; inverted repeat boundaries; nucleotide diversity; phylogenetic analysis; repetitive elements; selective pressure.
Conflict of interest statement
The authors declare that the research was conducted without any commercial or financial relationships that could potentially create conflicts of interest.
Figures




References
-
- Patchell M.J., Roalson E.H., Hall J.C. Resolved phylogeny of Cleomaceae based on all three genomes. Taxon. 2014;63:315–328. doi: 10.12705/632.17. - DOI
-
- Feodorova T.A., Voznesenskaya E.V., Edwards G.E., Roalson E.H. Biogeographic Patterns of Diversification and the Origins of C4 in Cleome (Cleomaceae) Syst. Bot. 2010;35:811–826. doi: 10.1600/036364410X539880. - DOI
-
- Iltis H.H. Studies in the Cleomaceae II: Cleome boliviensis, a new, spiny, large-flowered Andean species. Novon. 2005;15:146–155.
MeSH terms
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

