Cerulenin partially corrects the disrupted developmental transcriptomic signature in Huntington's disease striatal medium spiny neurons
- PMID: 40336225
- PMCID: PMC12277964
- DOI: 10.1093/stmcls/sxaf029
Cerulenin partially corrects the disrupted developmental transcriptomic signature in Huntington's disease striatal medium spiny neurons
Abstract
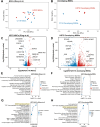
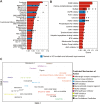
Huntington's disease (HD) is a neurodegenerative disorder caused by an expansion of CAG repeats in exon 1 of the huntingtin (HTT) gene, resulting in a mutant HTT (mHTT) protein. Although mHTT is expressed in all tissues, it significantly affects medium spiny neurons (MSNs) in the striatum, resulting in their loss and the subsequent motor function impairment in HD. While HD symptoms typically emerge in midlife, disrupted MSN neurodevelopment is important. To explore the effects of mHTT on MSN development, we differentiated HD-induced pluripotent stem cells (iPSCs) and isogenic controls into neuronal stem cells, and then generated a developing MSN population encompassing early, intermediate progenitors, and nascent MSNs. Single-cell RNA sequencing revealed that the developmental trajectory of MSNs in our model closely emulated the trajectory of human fetal striatal neurons. However, in the HD MSN cultures, several crucial genes required for proper MSN maturation were downregulated, including members of the DLX family of transcription factors. Our analysis also uncovered a progressive dysregulation of multiple HD-related pathways as MSNs developed, including the NRF2-mediated oxidative stress response and mitogen-activated protein kinase signaling. Using the transcriptional profile of developing HD MSNs, we searched the L1000 dataset for small molecules that induce the opposite gene expression pattern. We pinpointed numerous small molecules with known benefits in HD models and previously untested novel molecules. A top candidate, Cerulenin, partially restored the DARPP-32 levels and electrical activity in HD MSNs, and also modulated genes involved in multiple HD-related pathways.
Keywords: Huntington’s disease; development; induced pluripotent stem cells; medium spiny neurons; neurodegeneration; single-cell RNAseq.
© The Author(s) 2025. Published by Oxford University Press.
Conflict of interest statement
The authors have no financial/personal interest or belief that could affect their objectivity except AAG. No potential competing interests exist. AAG has financial interest in Image Analyst Software.
Figures






References
-
- Hedreen JC, Peyser CE, Folstein SE, Ross CA.. Neuronal loss in layers V and VI of cerebral cortex in Huntington’s disease. Neurosci Lett. 1991;133:257-261. https://doi.org/ 10.1016/0304-3940(91)90583-f - DOI - PubMed
-
- Cudkowicz M, Kowall NW.. Degeneration of pyramidal projection neurons in Huntington’s disease cortex. Ann Neurol. 1990;27:200-204. https://doi.org/ 10.1002/ana.410270217 - DOI - PubMed
-
- The Huntington’s Disease Collaborative Research Group. A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell. 1993;72:971-983. - PubMed
-
- Barnat M, Capizzi M, Aparicio E, et al. Huntington’s disease alters human neurodevelopment. Science. 2020;369:787-793. https://doi.org/ 10.1126/science.aax3338 - DOI - PMC - PubMed
-
- Molero AE, Gokhan S, Gonzalez S, et al. Impairment of developmental stem cell-mediated striatal neurogenesis and pluripotency genes in a knock-in model of Huntington’s disease. Proc Natl Acad Sci USA. 2009;106:21900-21905. https://doi.org/ 10.1073/pnas.0912171106 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

