Probing and manipulating the gut microbiome with chemistry and chemical tools
- PMID: 40336799
- PMCID: PMC12056425
- DOI: 10.1017/gmb.2025.4
Probing and manipulating the gut microbiome with chemistry and chemical tools
Abstract
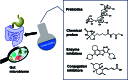
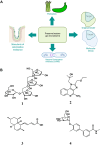
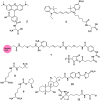
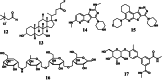
The human gut microbiome represents an extended "second genome" harbouring about 1015 microbes containing >100 times the number of genes as the host. States of health and disease are largely mediated by host-microbial metabolic interplay, and the microbiome composition also underlies the differential responses to chemotherapeutic agents between people. Chemical information will be the key to tackle this complexity and discover specific gut microbiome metabolism for creating more personalised interventions. Additionally, rising antibiotic resistance and growing awareness of gut microbiome effects are creating a need for non-microbicidal therapeutic interventions. We classify chemical interventions for the gut microbiome into categories like molecular decoys, bacterial conjugation inhibitors, colonisation resistance-stimulating molecules, "prebiotics" to promote the growth of beneficial microbes, and inhibitors of specific gut microbial enzymes. Moreover, small molecule probes, including click chemistry probes, artificial substrates for assaying gut bacterial enzymes and receptor agonists/antagonists, which engage host receptors interacting with the microbiome, are some other promising developments in the expanding chemical toolkit for probing and modulating the gut microbiome. This review explicitly excludes "biologics" such as probiotics, bacteriophages, and CRISPR to concentrate on chemistry and chemical tools like chemoproteomics in the gut-microbiome context.
Keywords: chemical probes; chemistry; conjugation inhibitors; gut microbiome; prebiotics.
© The Author(s) 2025.
Conflict of interest statement
The author Dr. Pavan Kumar Mantravadi is employee of Cytokinetics, South San Francisco, CA, USA. However, his company did not contribute to this written material. The remaining authors declare that the research/writing was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The paper was initiated by Dr. Anutthaman Parthasarathy and Dr. Ganesh Pandian Namasivayam, while the former was on an academic visit to Japan.
Figures
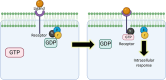
References
-
- Abt MC, Buffie CG, Sušac B, Becattini S, Carter RA, Leiner I, Keith JW, Artis D, Osborne LC and Pamer EG (2016) TLR-7 activation enhances IL-22-mediated colonization resistance against vancomycin-resistant enterococcus. Science Translational Medicine 8(327), 327ra325. 10.1126/scitranslmed.aad6663. - DOI - PMC - PubMed
-
- Adhikari AA, Seegar TCM, Ficarro SB, McCurry MD, Ramachandran D, Yao L, Chaudhari SN, Ndousse-Fetter S, Banks AS, Marto JA, et al. (2020) Development of a covalent inhibitor of gut bacterial bile salt hydrolases. Nature Chemical Biology 16(3), 318–326. 10.1038/s41589-020-0467-3. - DOI - PMC - PubMed
Publication types
LinkOut - more resources
Full Text Sources
