Analytical methods in studying cell force sensing: principles, current technologies and perspectives
- PMID: 40337625
- PMCID: PMC12057814
- DOI: 10.1093/rb/rbaf007
Analytical methods in studying cell force sensing: principles, current technologies and perspectives
Abstract
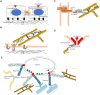
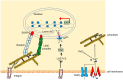
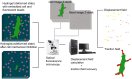
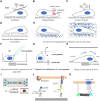
Mechanical stimulation plays a crucial role in numerous biological activities, including tissue development, regeneration and remodeling. Understanding how cells respond to their mechanical microenvironment is vital for investigating mechanotransduction with adequate spatial and temporal resolution. Cell force sensing-also known as mechanosensation or mechanotransduction-involves force transmission through the cytoskeleton and mechanochemical signaling. Insights into cell-extracellular matrix interactions and mechanotransduction are particularly relevant for guiding biomaterial design in tissue engineering. To establish a foundation for mechanical biomedicine, this review will provide a comprehensive overview of cell mechanotransduction mechanisms, including the structural components essential for effective mechanical responses, such as cytoskeletal elements, force-sensitive ion channels, membrane receptors and key signaling pathways. It will also discuss the clutch model in force transmission, the role of mechanotransduction in both physiology and pathological contexts, and biomechanics and biomaterial design. Additionally, we outline analytical approaches for characterizing forces at cellular and subcellular levels, discussing the advantages and limitations of each method to aid researchers in selecting appropriate techniques. Finally, we summarize recent advancements in cell force sensing and identify key challenges for future research. Overall, this review should contribute to biomedical engineering by supporting the design of biomaterials that integrate precise mechanical information.
Keywords: biomaterial design; biomaterial–cell interaction; biosensor; cell biomechanics.
© The Author(s) 2025. Published by Oxford University Press.
Figures






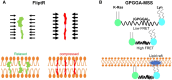
References
-
- He J, Liu Q, Zheng S, Shen R, Wang X, Gao J, Wang Q, Huang J, Ding J. Enlargement, reduction, and even reversal of relative migration speeds of endothelial and smooth muscle cells on biomaterials simply by adjusting RGD nanospacing. ACS Appl Mater Interfaces 2021;13:42344–56. - PubMed
-
- Liu Q, Zheng S, Ye K, He J, Shen Y, Cui S, Huang J, Gu Y, Ding J. Cell migration regulated by RGD nanospacing and enhanced under moderate cell adhesion on biomaterials. Biomaterials 2020;263:120327. - PubMed
-
- Haynie DT. Molecular physiology of the tensin brotherhood of integrin adaptor proteins. Proteins 2014;82:1113–27. - PubMed
Publication types
LinkOut - more resources
Full Text Sources

