The evaluation of antimicrobial resistance rates in infections caused by uropathogenic Escherichia coli strains collected from the south of Lebanon
- PMID: 40337688
- PMCID: PMC12053396
- DOI: 10.18502/ijm.v17i2.18386
The evaluation of antimicrobial resistance rates in infections caused by uropathogenic Escherichia coli strains collected from the south of Lebanon
Abstract
Background and objectives: Uropathogenic Escherichia coli (UPEC) is a leading cause of urinary tract infections, which are a significant public health concern worldwide. Antibiotic resistance among UPEC isolates is an increasing challenge, necessitating a better understanding of the resistance patterns and underlying genetic mechanisms. This study examined the prevalence of antibiotic resistance phenotypes and the detection of specific resistance genes among patients with UPEC infections in Sheikh Ragheb Harb University Hospital in south Lebanon.
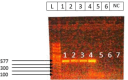
Materials and methods: Antimicrobial resistance phenotype of 104 urine samples was tested to determine the resistance percentages for various antibiotics including ampicillin, gentamicin, ciprofloxacin, tetracycline, bactrim, meropenem, and imipenem using disk diffusion test. Additionally, molecular analysis like polymerase chain reaction (PCR) was performed to detect the presence of bla SHV, qnrA, tetA, dfrA1, aac3, bla OXA and bla IMP resistance genes.
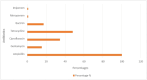
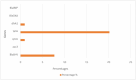
Results: The antimicrobial resistance testing revealed the following resistance percentages for various antibiotics: ampicillin (100%), gentamicin (15.38%), ciprofloxacin (34.61%), tetracycline (48.07%), bactrim (17.3%), meropenem (0.96%) and imipenem (0.96%). The analysis of resistance genes showed the presence of bla SHV (7.96%), qnrA (0.96%), tetA (20.19%), and dfrA1 (0.96%) genes, while the aac3, bla OXA, and bla IMP genes were not detected.
Conclusion: The high rates of antibiotic resistance observed, particularly to ampicillin and tetracycline, highlight the need for more judicious antibiotic use and the development of alternative treatment strategies to combat UPEC infections. These results can inform antimicrobial stewardship programs and guide the selection of appropriate empiric therapy for urinary tract infections.
Keywords: Antibiotic resistance; Polymerase chain reaction; Resistance genes; Urinary tract infection; Uropathogenic Escherichia coli.
Copyright © 2025 The Authors. Published by Tehran University of Medical Sciences.
Figures
References
LinkOut - more resources
Full Text Sources
