Diagnostic biomarkers and immune infiltration profiles common to COVID-19, acute myocardial infarction and acute ischaemic stroke using bioinformatics methods and machine learning
- PMID: 40340571
- PMCID: PMC12060493
- DOI: 10.1186/s12883-025-04212-6
Diagnostic biomarkers and immune infiltration profiles common to COVID-19, acute myocardial infarction and acute ischaemic stroke using bioinformatics methods and machine learning
Abstract
Background: COVID-19 is a disease that affects people globally. Beyond affecting the respiratory system, COVID-19 patients are at an elevated risk for both venous and arterial thrombosis. This heightened risk contributes to an increased probability of acute complications, including acute myocardial infarction (AMI) and acute ischemic stroke (AIS). Given the unclear relationship between COVID-19, AMI, and AIS, it is crucial to gain a deeper understanding of their associations and potential molecular mechanisms. This study aims to utilize bioinformatics to analyze gene expression data, identify potential therapeutic targets and biomarkers, and explore the role of immune cells in the disease.
Methods: This study employed three Gene Expression Omnibus (GEO) datasets for analysis, which included data on COVID-19, AMI and AIS. We performed enrichment analysis on the co-DEGs for these three diseases to clarify gene pathways and functions, and also examined the relationship between co-DEGs and immune infiltration. Machine learning techniques and protein-protein interaction networks (PPI) were used to identify hub genes within the co-DEGs. Finally, we employed a dual validation strategy integrating independent GEO datasets and in vitro experiments with human blood samples to comprehensively assess the reliability of our experimental findings.
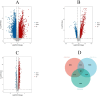
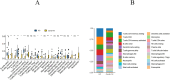
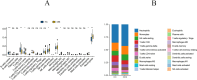
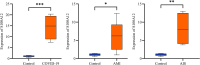
Results: We identified 88 co-DEGs associated with COVID-19, AMI and AIS. Enrichment analysis results indicated that co-DEGs were significantly enriched in immune inflammatory responses related to leukocytes and neutrophils. Immune infiltration analysis revealed significant differences in immune cell populations between the disease group and the normal group. Finally, genes selected through machine learning methods included: CLEC4E, S100A12, and IL1R2. Based on the PPI network, the top ten most influential DEGs were identified as MMP9, TLR2, TLR4, ITGAM, S100A12, FCGR1A, CD163, FCER1G, FPR2, and CLEC4D. The integration of the protein-protein interaction (PPI) network with machine learning techniques facilitated the identification of S100A12 as a potential common biomarker for early diagnosis and a therapeutic target for all three diseases. Ultimately, validation of S100A12 showed that it was consistent with our experimental results, confirming its reliability as a biomarker. Moreover, it demonstrated good diagnostic performance for the three diseases.
Conclusion: We employed bioinformatics methods and machine learning to investigate common diagnostic biomarkers and immune infiltration characteristics of COVID-19, AMI and AIS. Functional and pathway analyses indicated that the co-DEGs were primarily enriched in immune inflammatory responses related to leukocytes and neutrophils. Through two machine learning approaches and the PPI network, and subsequent validation and evaluation, we identified S100A12 as a potential common therapeutic target and biomarker related to immune response that may influence these three diseases.
Keywords: Bioinformatics; Biomarker; Blood clotting abnormality; COVID-19; Immune infiltration; Machine learning.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Data retrieved from the GEO database were uploaded in accordance with the guidelines established by the GEO Ethics, Law and Policy Group, so ethical review and informed consent were not required. The research involving human subjects has been approved by the Medical Research Ethics Review Committee of the General Hospital of Ningxia Medical University (Approval No.: KYLL-2025–0941). The study was conducted in compliance with local laws and institutional requirements, and informed consent was obtained from all participants. Consent for publication: All authors have given consent for publication. Competing interests: The authors declare no competing interests.
Figures












Similar articles
-
Identification and validation of diagnostic biomarkers of coronary artery disease progression in type 1 diabetes via integrated computational and bioinformatics strategies.Comput Biol Med. 2023 Jun;159:106940. doi: 10.1016/j.compbiomed.2023.106940. Epub 2023 Apr 15. Comput Biol Med. 2023. PMID: 37075605
-
Identification and analysis of key genes associated with acute myocardial infarction by integrated bioinformatics methods.Medicine (Baltimore). 2021 Apr 16;100(15):e25553. doi: 10.1097/MD.0000000000025553. Medicine (Baltimore). 2021. PMID: 33847684 Free PMC article.
-
Identification of diagnostic candidate genes in COVID-19 patients with sepsis.Immun Inflamm Dis. 2024 Oct;12(10):e70033. doi: 10.1002/iid3.70033. Immun Inflamm Dis. 2024. PMID: 39377750 Free PMC article.
-
Spatial Transcriptomics of the Respiratory System.Annu Rev Physiol. 2025 Feb;87(1):447-470. doi: 10.1146/annurev-physiol-022724-105144. Epub 2025 Feb 3. Annu Rev Physiol. 2025. PMID: 39353142 Review.
-
The Role of Infection and Inflammation in the Pathogenesis of Pediatric Arterial Ischemic Stroke.Semin Pediatr Neurol. 2022 Dec;44:100995. doi: 10.1016/j.spen.2022.100995. Epub 2022 Sep 10. Semin Pediatr Neurol. 2022. PMID: 36456035 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

