Whole-genome sequencing analysis to identify antimicrobial resistance regions and virulence factors in Mycobacterium tuberculosis isolates from the Amhara Region, Ethiopia
- PMID: 40341578
- PMCID: PMC12062464
- DOI: 10.1038/s41598-025-01241-6
Whole-genome sequencing analysis to identify antimicrobial resistance regions and virulence factors in Mycobacterium tuberculosis isolates from the Amhara Region, Ethiopia
Abstract
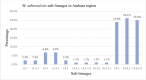
Tuberculosis caused by Mycobacterium tuberculosis complex is a significant global health burden, with drug-resistant TB, especially multidrug-resistant TB, causing severe challenges to treatment. In Ethiopia, a high TB-burden country, drug resistance has continued spreading. However, some studies indicate genetic diversity, transmission dynamics, and resistance-conferring mutations by using targeted amplification, there are limited reports of whole genome sequencing analysis to uncover the antimicrobial resistance and virulent genes. Based on that, the objective of this project was to identify antimicrobial resistance regions and characterize virulence factors in M. tuberculosis isolates through in silico whole-genome sequence analysis. A FASTQ file of 45 M. tuberculosis isolates whole genome sequence was downloaded from the SAR database. Following quality control using FASTQC coupled with MultiQC and trimming with Trimmomatic, de novo assembly was conducted using SPAdes. The Burrows-Wheeler Aligner was used for mapping against the M. tuberculosis H37Rv reference genome, followed by variant calling with FreeBayes. In silico spoligotyping was performed using SpoTyping, and drug resistance mutations were identified with TB-Profiler and validated using Mykrobe. Virulence factors were detected through ABRicate and the Virulence Factor Database. STRING was used to network the virulent genes. All statistical analyses were performed using R software. This study revealed the most prevalent TB-lineage in the Amhara region was L4 (58.53%), followed by L3 (34.15%), and L1 (4.88%), and in silico spoligotyping classified 90.24% of the isolates into 12 shared types, with SIT 149 (41.46%) and SIT 21 (14.63%) as the most frequent spoligotypes. Seven major genotypic families were identified, with T3-ETH being the dominant family (48.78%). Drug resistance analysis revealed that 38 isolates (92.7%) were multidrug-resistant, and 1 (2.4%) was pre-extensively drug-resistant. Lineage 4 (59%) and its sub-lineage 4.2.2 (51.3%) show the highest resistance. The most frequent mutations to rifampicin, isoniazid, pyrazinamide, ethambutol, streptomycin, ethionamide, fluoroquinolone, and 2nd-line injectable drugs occurred at rpoB Ser450Leu, katG Ser315Thr, pncA c.-11A > G, embB Gly406Ala, rpsL Lys43Arg, Lys88Thr, ethA Met1, gyrA Ala90Val, Asp94Asn, and rrs 1401A > G, respectively. Additionally, a mutation at the mmpR5 gene for bedaquiline and clofazimine resistance occurred in one isolate. A total of 67 virulence genes were identified and 63 of them occurred in all isolates. The high prevalence of MDR-TB and the detection of resistance to both first- and second-line drugs in this study underscore the urgent need for enhanced TB control measures in the Amhara region.
Keywords: Mycobacterium tuberculosis; Drug resistance; Virulence; Whole genome and Amhara.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests. Ethics statement: Not applicable as there is not any animal or human subject involved directly in the study. The study utilized publicly available raw WGS data. Demographic and clinical data of patients were accessed from an open-source database.
Figures
References
-
- Lin, S. Y. G. & Desmond, E. P. Molecular diagnosis of tuberculosis and drug resistance. Clin. Lab. Med.34(2), 297–314 (2014). - PubMed
-
- WHO. Global tuberculosis report 2024. Geneva: World Health Organization; 2024. Licence: CC BY-NC-SA 3.0 IGO [Internet]. https://www.who.int/teams/global-tuberculosis-programme/tb-reports/globa... (2024).
-
- WHO Africa. Country Disease Outlook Ethiopia, 55–58. https://www.afro.who.int/sites/default/files/2023-08/Ethiopia.pdf (2023).
-
- US Agency for International Development. Ethiopia tuberculosis roadmap overview, Fiscal Year 2023. https://www.usaid.gov/sites/default/files/2024-02/Ethiopia_TB_roadmap_na... (2023).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous