The telomere-to-telomere genome of Pucai () (Typha angustifolia L.): a distinctive semiaquatic vegetable with lignin and chlorophyll as quality characteristics
- PMID: 40343350
- PMCID: PMC12058305
- DOI: 10.1093/hr/uhaf079
The telomere-to-telomere genome of Pucai () (Typha angustifolia L.): a distinctive semiaquatic vegetable with lignin and chlorophyll as quality characteristics
Abstract
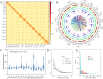
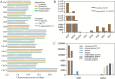
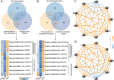
Pucai () (Typha angustifolia L.), within the Typha spp., is a distinctive semiaquatic vegetable. Lignin and chlorophyll are two crucial traits and quality indicators for Pucai. In this study, we assembled a 207.00-Mb high-quality gapless genome of Pucai, telomere-to-telomere (T2T) level with a contig N50 length of 13.73 Mb. The most abundant type of repetitive sequence, comprising 16.98% of the genome, is the long terminal repeat retrotransposons (LTR-RT). A total of 30 telomeres and 15 centromeric regions were predicted. Gene families related to lignin, chlorophyll biosynthesis, and disease resistance were greatly expanded, which played important roles in the adaptation of Pucai to wetlands. The slow evolution of Pucai was indicated by the σ whole-genome duplication (WGD)-associated Ks peaks from different Poales and the low activity of recent LTR-RT in Pucai. Meanwhile, we found a unique WGD event in Typhaceae. A statistical analysis and annotation of genomic variations were conducted in interspecies and intraspecies of Typha. Based on the T2T genome, we constructed lignin and chlorophyll metabolic pathways of Pucai. Subsequently, the candidate structural genes and transcription factors that regulate lignin and chlorophyll biosynthesis were identified. The T2T genomic resources will provide molecular information for lignin and chlorophyll accumulation and help to understand genome evolution in Pucai.
© The Author(s) 2025. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures





References
-
- Claudia C, Heather K, Jeffrey RR. et al. Intercontinental dispersal of Typha angustifolia and T. latifolia between Europe and North America has implications for Typha invasions. Biol Invasions. 15:1377–90
-
- Smith SG. Experimental and natural hybrids in North American Typha (Typhaceae). Am Midl Nat. 1967;78:257–87
-
- Smith SG. Typha: its taxonomy and the ecological significance of hybrids. Arch Hydrobiol. 1987;27:129–38
LinkOut - more resources
Full Text Sources
Miscellaneous

