CILP1 interacting with YBX1 promotes hypertrophic scar formation by suppressing PPARs transcription
- PMID: 40346063
- PMCID: PMC12064789
- DOI: 10.1038/s41419-025-07554-8
CILP1 interacting with YBX1 promotes hypertrophic scar formation by suppressing PPARs transcription
Abstract
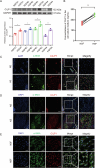
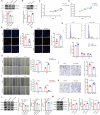
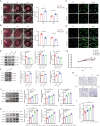
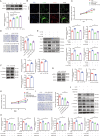
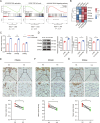
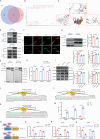
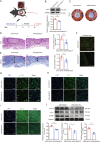
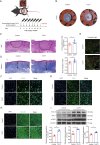
Hypertrophic scar (HS) represents the most prevalent form of skin fibrosis, significantly impacting the quality of life. Despite this, the molecular mechanisms driving HS formation remain largely undefined, impeding the development of effective treatments. The study showed that Cartilage Intermediate Layer Protein 1 (CILP1) was predominantly expressed in myofibroblasts and was up-regulated in various forms of skin fibrosis, including human hypertrophic and keloid scars, and in animal models of HS. Notably, we detected elevated serum levels of CILP1 in fifty-two patients with HS compared to twenty healthy individuals, suggesting its potential as a novel biomarker. The findings indicated that CILP1 was involved in a negative feedback loop with TGF-β and inhibited the transcription of Peroxisome Proliferator-Activated Receptors (PPARs) via interaction with Y-box-binding protein 1 (YBX1). This interaction promoted cell proliferation, migration, and collagen production in hypertrophic scar fibroblasts (HSFs). In vivo studies further confirmed that CILP1 knockdown markedly reduced HS formation, whereas administration of recombinant human CILP1 protein exacerbated it. These discoveries illuminated the CILP1-YBX1-PPARs signaling pathway as a key regulator of HS formation, offering a foundation for novel therapeutic approaches.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests. Ethical approval and consent to participate: The study was conducted in accordance with the principles of the Helsinki Declaration. All patients were informed of the study purpose and processes and provided the informed consent for participation into the present work. The Ethics Committee of Xijing Hospital affiliated to the Fourth Military Medical University approved the present work (approval number: No.KY20203134-1). Our animal experiments followed the Guide for the Care and Use of Laboratory Animals. Experimental protocols were approved by the Committee on the Ethics of Animal Experiments of Fourth Military Medical University; approval number: No.20230033. Consent for publication: All authors have read and approved the final manuscript.
Figures


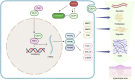
References
-
- Gold MH, Berman B, Clementoni MT, Gauglitz GG, Nahai F, Murcia C. Updated international clinical recommendations on scar management: part 1-evaluating the evidence. Dermatol Surg. 2014;40:817–24. - PubMed
MeSH terms
Substances
Grants and funding
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
- 82072182/National Natural Science Foundation of China (National Science Foundation of China)
- 82372530/National Natural Science Foundation of China (National Science Foundation of China)
LinkOut - more resources
Full Text Sources
Research Materials

