Parent-of-origin regulation by maternal auts2 shapes neurodevelopment and behavior in fish
- PMID: 40346605
- PMCID: PMC12063280
- DOI: 10.1186/s13059-025-03600-y
Parent-of-origin regulation by maternal auts2 shapes neurodevelopment and behavior in fish
Abstract
Background: Parental experience can influence progeny behavior through gamete-mediated non-genetic inheritance, that is, mechanisms that do not involve changes in inherited DNA sequence. However, underlying mechanisms remain poorly understood in vertebrates, especially for maternal effects. Here, we use the medaka, a model fish species, to investigate the role of auts2a, the ortholog of human AUTS2, a gene repressed in the fish oocyte following maternal stress and associated with neurodevelopmental disorders.
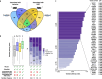
Results: We show that auts2a expression in the oocyte influences long-term progeny behavior, including anxiety-like behavior and environment recognition capabilities. Using single-nuclei RNA-sequencing, we reveal that maternal auts2a influences gene expression in neural cell populations during neurodevelopment. We also show that maternal auts2a knock-out triggers differences in maternally inherited factors, including early embryonic transcriptional and post-transcriptional regulators.
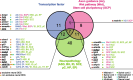
Conclusions: Together, our results reveal the unsuspected role of an autism-related gene expressed in the mother's oocyte in shaping progeny neurodevelopment and behavior. Finally, we report that auts2a/AUTS2 is part of a group of evolutionarily conserved genes associated with human neurodevelopmental disorders and expressed in oocytes across species, from fish to mammals. These findings raise important questions about their potential role in the non-genetic regulation of progeny neurodevelopment and behavior in vertebrates.
Keywords: ASD; Intergenerational effect; Medaka; Neurodevelopmental disorder; Non-genetic inheritance.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests.
Figures




Similar articles
-
auts2 Features and Expression Are Highly Conserved during Evolution Despite Different Evolutionary Fates Following Whole Genome Duplication.Cells. 2022 Aug 30;11(17):2694. doi: 10.3390/cells11172694. Cells. 2022. PMID: 36078102 Free PMC article.
-
Bucky ball induces primordial germ cell increase in medaka.Gene. 2021 Feb 5;768:145317. doi: 10.1016/j.gene.2020.145317. Epub 2020 Nov 19. Gene. 2021. PMID: 33221537
-
Maternal temperature exposure impairs emotional and cognitive responses and triggers dysregulation of neurodevelopment genes in fish.PeerJ. 2019 Jan 31;7:e6338. doi: 10.7717/peerj.6338. eCollection 2019. PeerJ. 2019. PMID: 30723624 Free PMC article.
-
AUTS2 Gene: Keys to Understanding the Pathogenesis of Neurodevelopmental Disorders.Cells. 2021 Dec 21;11(1):11. doi: 10.3390/cells11010011. Cells. 2021. PMID: 35011572 Free PMC article. Review.
-
Untangle the Multi-Facet Functions of Auts2 as an Entry Point to Understand Neurodevelopmental Disorders.Front Psychiatry. 2021 Apr 23;12:580433. doi: 10.3389/fpsyt.2021.580433. eCollection 2021. Front Psychiatry. 2021. PMID: 33967843 Free PMC article. Review.
References
-
- Wehmer F, Porter RH, Scales B. Pre-mating and pregnancy stress in rats affects behaviour of grandpups. Nature. 1970;227:622. - PubMed
-
- Adrian-Kalchhauser I, Sultan SE, Shama LNS, Spence-Jones H, Tiso S, Keller Valsecchi CI, et al. Understanding “Non-genetic” Inheritance: Insights from Molecular-Evolutionary Crosstalk. Trends Ecol Evol. 2020;35:1078–89. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

