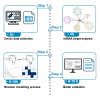
Protocol to identify regulatory modules in Parkinson's disease progression using miRNA data and Boolean modeling
- PMID: 40347476
- PMCID: PMC12136812
- DOI: 10.1016/j.xpro.2025.103769
Protocol to identify regulatory modules in Parkinson's disease progression using miRNA data and Boolean modeling
Abstract
Regulatory modules are molecules that interact functionally, driving disease processes. Here, we present a protocol for identifying regulatory modules in Parkinson's disease (PD) using cohort-specific microRNA (miRNA) data and Boolean modeling. We describe steps for omics data collection, biomolecule and miRNA target analysis, and Boolean model construction and simulation. We then detail procedures for validation of the model and results. The modules identified using this protocol explain how miRNA-driven mechanisms influence PD progression in disease cohorts. For complete details on the use and execution of this protocol, please refer to Hemedan et al.1.
Keywords: Bioinformatics; Molecular Biology; Neuroscience; Systems biology.
Copyright © 2025 The Author(s). Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
References
-
- Marek K., Chowdhury S., Siderowf A., Lasch S., Coffey C.S., Caspell-Garcia C., Simuni T., Jennings D., Tanner C.M., Trojanowski J.Q., et al. The Parkinson’s progression markers initiative (PPMI) – establishing a PD biomarker cohort. Ann. Clin. Transl. Neurol. 2018;5:1460–1477. doi: 10.1002/acn3.644. - DOI - PMC - PubMed
-
- Funahashi A. In: Introduction to Systems Biology. David O’Connor, editor. Humana Press; 2007. CellDesigner: A Graphical Biological Network Editor and Workbench Interfacing Simulator; pp. 422–434.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical