SEMTWIST Quantification of Biofilm Infection in Human Chronic Wound Using Scanning Electron Microscopy and Machine Learning
- PMID: 40358506
- PMCID: PMC12359142
- DOI: 10.1089/wound.2024.0291
SEMTWIST Quantification of Biofilm Infection in Human Chronic Wound Using Scanning Electron Microscopy and Machine Learning
Abstract
Objective: To develop scanning electron microscopy-based Trainable Weka (Waikato Environment for Knowledge Analysis) Intelligent Segmentation Technology (SEMTWIST), an open-source software tool, for structural detection and rigorous quantification of wound biofilm aggregates in complex human wound tissue matrix. Approach: SEMTWIST model was standardized to quantify biofilm infection (BFI) abundance in 240 distinct SEM images from 60 human chronic wound-edge biospecimens (four technical replicates of each specimen). Results from SEMTWIST were compared against human expert assessments and the gold standard for molecular BFI detection, that is, peptide nucleic acid fluorescence in situ hybridization (PNA-FISH). Results: Correlation and Bland-Altman plot demonstrated a robust correlation (r = 0.82, p < 0.01), with a mean bias of 1.25, and 95% limit of agreement ranging from -43.40 to 47.11, between SEMTWIST result and the average scores assigned by trained human experts. While interexpert variability highlighted potential bias in manual assessments, SEMTWIST provided consistent results. Bacterial culture detected infection but not biofilm aggregates. Whereas the wheat germ agglutinin staining exhibited nonspecific staining of host tissue components and failed to provide a specific identification of BFI. The molecular identification of biofilm aggregates using PNA-FISH was comparable with SEMTWIST, highlighting the robustness of the developed approach. Innovation: This study introduces a novel approach "SEMTWIST" for in-depth analysis and precise differentiation of biofilm aggregates from host tissue elements, enabling accurate quantification of BFI in chronic wound SEM images. Conclusion: Open-source SEMTWIST offers a reliable and robust framework for standardized quantification of BFI burden in human chronic wound-edge tissues, supporting clinical diagnosis and guiding treatment.
Keywords: biofilm infection; culture; human chronic wound; machine learning; scanning electron microscopy; wheat germ agglutinin.
Conflict of interest statement
AUTHOR DISCLOSURE AND GHOST WRITING
The authors declare no financial interests or conflicts of interest to disclose.
Figures
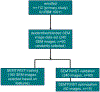





References
-
- Bjarnsholt T. The role of bacterial biofilms in chronic infections. APMIS 2013;121(136):1–51. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

