Dual fragmentation via collision-induced and oxygen attachment dissociations using water and its radicals for C=C position-resolved lipidomics
- PMID: 40360765
- PMCID: PMC12075507
- DOI: 10.1038/s42004-025-01525-y
Dual fragmentation via collision-induced and oxygen attachment dissociations using water and its radicals for C=C position-resolved lipidomics
Abstract
Oxygen attachment dissociation (OAD) is a tandem mass spectrometry (MS/MS) technique for annotating the positions of double bonds (C=C) in complex lipids. Although OAD has been used for untargeted lipidomics, its availability has been limited to the positive ion mode, requiring the independent use of a collision-induced dissociation (CID) method. In this study, we demonstrated the OAD MS/MS technique in the negative-ion mode for profiling phosphatidylserines, phosphatidylglycerols, phosphatidylinositols, and sulfatides, where the fragmentation mechanism remained consistent with that in the positive ion mode. Furthermore, we proposed optimal conditions for the simultaneous acquisition of CID- and OAD-specific fragment ions, termed OAciD, where oxygen atoms and hydroxy radicals facilitate C=C position-specific fragmentation, while residual water vapor induces cleavage of low-energy covalent bonds as observed in CID. Finally, theoretical fragment ions were implemented in MS-DIAL 5 to accelerate C=C position-resolved untargeted lipidomics. The OAciD methodology was used to illuminate brain region-specific marmoset lipidomes with C=C positional information, including the estimation of C=C positional isomer ratios. We also characterized the profiles of polyunsaturated fatty acid-containing lipids, finding that lipids containing omega-3 fatty acids were enriched in the cerebellum, whereas those containing omega-6 fatty acids were more abundant in the hippocampus and frontal lobe.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: M.O. and H. Takahashi are research scientists at Shimadzu, Inc., Japan. All the other authors declare that they have no competing interests.
Figures



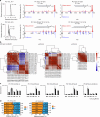
References
Grants and funding
- JP25wm0325071/Japan Agency for Medical Research and Development (AMED)
- JPMJND2305/MEXT | JST | National Bioscience Database Center (NBDC)
- JPMJER2101/MEXT | JST | Exploratory Research for Advanced Technology (ERATO)
- JPMJFR230H/MEXT | Japan Science and Technology Agency (JST)
- 24H00043/MEXT | Japan Society for the Promotion of Science (JSPS)
LinkOut - more resources
Full Text Sources

