Structure, function and assembly of soybean primary cell wall cellulose synthases
- PMID: 40365874
- PMCID: PMC12077881
- DOI: 10.7554/eLife.96704
Structure, function and assembly of soybean primary cell wall cellulose synthases
Abstract
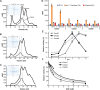
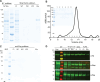
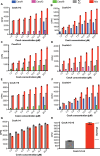
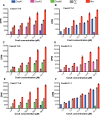
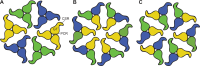
Plant cell walls contain a meshwork of cellulose fibers embedded into a matrix of other carbohydrate and non-carbohydrate-based biopolymers. This composite material exhibits extraordinary properties, from stretchable and pliable cell boundaries to solid protective shells. Cellulose, a linear glucose polymer, is synthesized and secreted across the plasma membrane by cellulose synthase (CesA), of which plants express multiple isoforms. Different subsets of CesA isoforms are necessary for primary and secondary cell wall biogenesis. Here, we structurally and functionally characterize the Glycine max (soybean) primary cell wall CesAs CesA1, CesA3, and CesA6. The CesA isoforms exhibit robust in vitro catalytic activity. Cryo-electron microscopy analyses reveal their assembly into homotrimeric complexes in vitro in which each CesA protomer forms a cellulose-conducting transmembrane channel with a large lateral opening. Biochemical and co-purification analyses demonstrate that different CesA isoforms interact in vitro, leading to synergistic cellulose biosynthesis. Interactions between CesA trimers are only observed between different CesA isoforms and require the class-specific region (CSR). The CSR forms a hook-shaped extension of CesA's catalytic domain at the cytosolic water-lipid interface. Negative stain and cryo-electron microscopy analyses of mixtures of different CesA isoform trimers reveal their side-by-side arrangement into loose clusters. Our data suggest a model by which CesA homotrimers of different isoforms assemble into cellulose synthase complexes to synthesize and secrete multiple cellulose chains for microfibril formation. Inter-trimer interactions are mediated by fuzzy interactions between their CSR extensions.
Keywords: cellulose; electron microscopy; intrinsically disordered domain; microfibril; molecular biophysics; rosette; structural biology.
© 2024, Ho, Pallinti et al.
Conflict of interest statement
RH, PP, LW, YW, JZ No competing interests declared
Figures









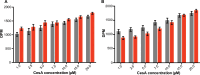

Update of
-
Structure, function and assembly of soybean primary cell wall cellulose synthases.bioRxiv [Preprint]. 2025 Jan 14:2024.02.13.580128. doi: 10.1101/2024.02.13.580128. bioRxiv. 2025. Update in: Elife. 2025 May 14;13:RP96704. doi: 10.7554/eLife.96704. PMID: 38405885 Free PMC article. Updated. Preprint.
Similar articles
-
Structure, function and assembly of soybean primary cell wall cellulose synthases.bioRxiv [Preprint]. 2025 Jan 14:2024.02.13.580128. doi: 10.1101/2024.02.13.580128. bioRxiv. 2025. Update in: Elife. 2025 May 14;13:RP96704. doi: 10.7554/eLife.96704. PMID: 38405885 Free PMC article. Updated. Preprint.
-
The molecular basis of plant cellulose synthase complex organisation and assembly.Biochem Soc Trans. 2021 Feb 26;49(1):379-391. doi: 10.1042/BST20200697. Biochem Soc Trans. 2021. PMID: 33616627 Review.
-
A Structural Study of CESA1 Catalytic Domain of Arabidopsis Cellulose Synthesis Complex: Evidence for CESA Trimers.Plant Physiol. 2016 Jan;170(1):123-35. doi: 10.1104/pp.15.01356. Epub 2015 Nov 10. Plant Physiol. 2016. PMID: 26556795 Free PMC article.
-
Features of the primary wall CESA complex in wild type and cellulose-deficient mutants of Arabidopsis thaliana.J Exp Bot. 2008;59(10):2627-37. doi: 10.1093/jxb/ern125. Epub 2008 May 20. J Exp Bot. 2008. PMID: 18495638 Free PMC article.
-
Updating Insights into the Catalytic Domain Properties of Plant Cellulose synthase (CesA) and Cellulose synthase-like (Csl) Proteins.Molecules. 2021 Jul 17;26(14):4335. doi: 10.3390/molecules26144335. Molecules. 2021. PMID: 34299608 Free PMC article. Review.
Cited by
-
Empirical evidence that glucan-interacting amino acid side chains within the transmembrane channel collectively facilitate cellulose synthase function.Plant Mol Biol. 2025 Jul 9;115(4):85. doi: 10.1007/s11103-025-01615-4. Plant Mol Biol. 2025. PMID: 40632364 Free PMC article.
References
-
- Afonine PV, Grosse-Kunstleve RW, Echols N, Headd JJ, Moriarty NW, Mustyakimov M, Terwilliger TC, Urzhumtsev A, Zwart PH, Adams PD. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallographica. Section D, Biological Crystallography. 2012;68:352–367. doi: 10.1107/S0907444912001308. - DOI - PMC - PubMed
-
- Baytshtok V, Chen JJ, Glynn SE, Nager AR, Grant RA, Baker TA, Sauer RT. Covalently linked HslU hexamers support a probabilistic mechanism that links ATP hydrolysis to protein unfolding and translocation. The Journal of Biological Chemistry. 2017;292:5695–5704. doi: 10.1074/jbc.M116.768978. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

