A bloom of a single bacterium shapes the microbiome during outdoor diatom cultivation collapse
- PMID: 40366134
- PMCID: PMC12172471
- DOI: 10.1128/msystems.00375-25
A bloom of a single bacterium shapes the microbiome during outdoor diatom cultivation collapse
Abstract
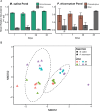
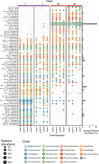
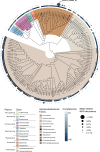
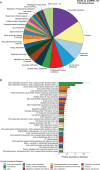
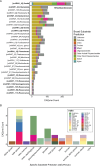
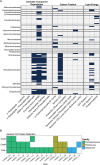
Algae-dominated ecosystems are fundamentally influenced by their microbiome. We lack information on the identity and function of bacteria that specialize in consuming algal-derived dissolved organic matter in high algal density ecosystems such as outdoor algal ponds used for biofuel production. Here, we describe the metagenomic and metaproteomic signatures of a single bacterial strain that bloomed during a population-wide crash of the diatom, Phaeodactylum tricornutum, grown in outdoor ponds. 16S rRNA gene data indicated that a single Kordia sp. strain (family Flavobacteriaceae) contributed up to 93% of the bacterial community during P. tricornutum demise. Kordia sp. expressed proteins linked to microbial antagonism and biopolymer breakdown, which likely contributed to its dominance over other microbial taxa during diatom demise. Analysis of accompanying downstream microbiota (primarily of the Rhodobacteraceae family) provided evidence that cross-feeding may be a pathway supporting microbial diversity during diatom demise. In situ and laboratory data with a different strain suggested that Kordia was a primary degrader of biopolymers during algal demise, and co-occurring Rhodobacteraceae exploited degradation molecules for carbon. An analysis of 30 Rhodobacteraceae metagenome assembled genomes suggested that algal pond Rhodobacteraceae commonly harbored pathways to use diverse carbon and energy sources, including carbon monoxide, which may have contributed to the prevalence of this taxonomic group within the ponds. These observations further constrain the roles of functionally distinct heterotrophic bacteria in algal microbiomes, demonstrating how a single dominant bacterium, specialized in processing senescing or dead algal biomass, shapes the microbial community of outdoor algal biofuel ponds.IMPORTANCEAquatic biogeochemical cycles are dictated by the activity of diverse microbes inhabiting the algal microbiome. Outdoor biofuel ponds provide a setting analogous to aquatic algal blooms, where monocultures of fast-growing algae reach high cellular densities. Information on the microbial ecology of this setting is lacking, and so we employed metagenomics and metaproteomics to understand the metabolic roles of bacteria present within four replicated outdoor ponds inoculated with the diatom Phaeodactylum tricornutum. Unexpectedly, after 29 days of cultivation, all four ponds crashed concurrently with a "bloom" of a single taxon assigned to the Kordia bacterial genus. We assessed how this dominant taxon influenced the chemical and microbial fate of the ponds following the crash, with the hypothesis that it was primarily responsible for processing senescent/dead algal biomass and providing the surrounding microbiota with carbon. Overall, these findings provide insight into the roles of microbes specialized in processing algal organic matter and enhance our understanding of biofuel pond microbial ecology.
Keywords: Kordia; biofuel ponds; cross-feeding; metagenomics; metaproteomics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

References
-
- Cole JJ. 1982. Interactions between bacteria and algae in aquatic ecosystems. Annu Rev Ecol Syst 13:291–314. doi: 10.1146/annurev.es.13.110182.001451 - DOI
-
- Wheeler PA, Kirchman DL. 1986. Utilization of inorganic and organic nitrogen by bacteria in marine systems. Limnol Oceanogr 31:998–1009. doi: 10.4319/lo.1986.31.5.0998 - DOI
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

