Calcium signaling controls early stage biofilm formation and dispersal in Vibrio fischeri
- PMID: 40366159
- PMCID: PMC12186489
- DOI: 10.1128/jb.00077-25
Calcium signaling controls early stage biofilm formation and dispersal in Vibrio fischeri
Abstract
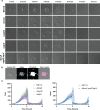
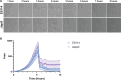
Bacterial dispersal from a biofilm is presently the least-studied step of the biofilm life cycle. The symbiotic bacterial species Vibrio fischeri is a model organism for studying biofilms relevant to a eukaryotic host; however, methodology is lacking to readily study the dispersal of this microbe from biofilms formed in the lab. Here, we adapted a time-lapse assay to visualize biofilm dispersal by V. fischeri. We observed biofilm formation and dispersal for multiple V. fischeri isolates, which displayed a variety of biofilm architecture phenotypes and dispersal dynamics. We then investigated V. fischeri strain ES114 using genetic tools and mutants available for this strain. ES114 exhibited calcium-dependent biofilm formation followed by a rapid (less than 10 min) coordinated dispersal event that occurred approximately 5 h from the experimental start. Biofilm dispersal was largely independent of the dispersal-promoting protease encoded by lapG. Although we found no role under our conditions for either biofilm formation or dispersal for several other factors including polysaccharides and autoinducers, we determined that biofilm formation was enhanced, and dispersal was delayed, with increased concentrations of calcium. Furthermore, biofilm formation depended on the calcium-responsive diguanylate cyclase (DGC) CasA, and dispersal could be modulated by overexpressing CasA. Our work has thus developed a new tool for the V. fischeri field and uncovered a key role for calcium signaling and c-di-GMP in early biofilm formation and dispersal in V. fischeri.
Importance: Biofilm formation and dispersal are critical steps in both symbiotic and pathogenic colonization. Relative to biofilm formation, the process of dispersal in the model symbiont Vibrio fischeri, and other bacteria, is understudied. Here, we adapted an imaging assay to study early biofilm formation and the dispersal process in V. fischeri. We demonstrated that our assay can quantify biofilm formation and dispersal over time, can reveal phenotypic differences in diverse natural wild-type isolates, and is sensitive enough to investigate the impact of environmental factors. Our data confirm that calcium is a potent biofilm formation signal and identify the diguanylate cyclase CasA as a key regulator. This work leads the way for more in-depth research about unknown mechanisms of biofilm dispersal.
Keywords: Vibrio fischeri; biofilms; dispersal; quorum sensing; video microscopy.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

