Hydrogen-dependent dissimilatory nitrate reduction to ammonium enables growth of Campylobacterota isolates
- PMID: 40367351
- PMCID: PMC12286921
- DOI: 10.1093/ismejo/wraf092
Hydrogen-dependent dissimilatory nitrate reduction to ammonium enables growth of Campylobacterota isolates
Abstract
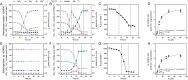
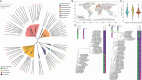
Dissimilatory nitrate reduction to ammonium (DNRA) is a key process used by diverse microorganisms in the global nitrogen cycle. For long, DNRA has been considered primarily as an organotrophic reaction, despite evidence that oxidation of inorganic electron donors also supports DNRA. Evidence of DNRA coupling with molecular hydrogen (H2) oxidation has been reported for several microbial isolates; however, the underlying physiology of the microbial process remains understudied. In this study, we report the isolation of two Campylobacterota strains, Aliarcobacter butzleri hDNRA1 and Sulfurospirillum sp. hDNRA2, which grow using H2 as the sole electron donor for DNRA, and physiological insights gained from a close examination of hydrogenotrophic DNRA in these isolates. In both batch and continuous cultures, DNRA sensu stricto (i.e. NO3- reduction that includes stoichiometric NO2--to-NH4+ reduction) was strictly dependent on the presence of H2 and exhibited stoichiometric coupling with H2 oxidation, indicating that electrons required for NO2- reduction were unequivocally derived from H2. Successful chemostat incubation further demonstrated that hydrogenotrophic DNRA is viable under NO3--limiting, H2-excess conditions. Genomic and transcriptomic analyses identified group 1b [NiFe]-hydrogenase and cytochrome c552 nitrite reductase as the key enzymes catalyzing hydrogenotrophic DNRA. In addition, metagenomic surveys revealed that bacteria capable of hydrogenotrophic DNRA are taxonomically diverse and abundant in various ecosystems, particularly in the vicinity of deep-sea hydrothermal vents. These findings, integrating physiological, genomic, and transcriptomic analyses, clarify that H2 can solely serve as a growth-supporting electron donor for DNRA and suggest potential significance of this microbial process in nitrogen- and hydrogen-related environmental biogeochemical cycles.
Keywords: Campylobacterota; dissimilatory nitrate reduction to ammonium (DNRA); hydrogenases; nitrogen cycling; respiratory hydrogen oxidation.
© The Author(s) 2025. Published by Oxford University Press on behalf of the International Society for Microbial Ecology.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Smil V. Enriching the Earth: Fritz Haber, Carl Bosch, and the Transformation of World Food Production. Cambridge: The MIT Press, 2000. 10.7551/mitpress/2767.001.0001 - DOI
MeSH terms
Substances
Grants and funding
- DP210101595/Australian Research Council (ARC) Discovery Project
- APP1178715/National Health and Medical Research Council (NHMRC) EL2 Fellowship
- RS-2024-00341771/National Research Foundation of Korea
- NRF-2022R1A4A5031447/National Research Foundation of Korea
- NRF-2021R1C1C1008303/National Research Foundation of Korea
LinkOut - more resources
Full Text Sources

