Identification of undetected SARS-CoV-2 infections by clustering of Nucleocapsid antibody trajectories
- PMID: 40368905
- PMCID: PMC12078723
- DOI: 10.1038/s41467-025-57370-z
Identification of undetected SARS-CoV-2 infections by clustering of Nucleocapsid antibody trajectories
Abstract
During the COVID-19 pandemic, numerous SARS-CoV-2 infections remained undetected. We combined results from routine monthly nose and throat swabs, and self-reported positive swab tests, from a UK household survey, linked to national swab testing programme data from England and Wales, together with Nucleocapsid (N-)antibody trajectories clustered using a longitudinal variation of K-means (N = 185,646) to estimate the number of infections undetected by either approach. Using N-antibody (hypothetical) infections and swab-positivity, we estimated that 7.4% (95%CI: 7.0-7.8%) of all true infections (detected and undetected) were undetected by both approaches, 25.8% (25.5-26.1%) by swab-positivity-only and 28.6% (28.4-28.9%) by trajectory-based N-antibody-classifications-only. Congruence with swab-positivity was respectively much poorer and slightly better with N-antibody classifications based on fixed thresholds or fourfold increases. Using multivariable logistic regression N-antibody seroconversion was more likely as age increased between 30-60 years, in non-white participants, those less (recently/frequently) vaccinated, for lower cycle threshold values in the range above 30, and in symptomatic and Delta (vs. BA.1) infections. Comparing swab-positivity data sources showed that routine monthly swabs were insufficient to detect infections and incorporating national testing programme/self-reported data substantially increased detection. Overall, whilst N-antibody serosurveillance can identify infections undetected by swab-positivity, optimal use requires fourfold-increase-based or trajectory-based analysis.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: This study was funded by the UK Health Security Agency and the Department of Health and Social Care with in-kind support from the Welsh Government, the Department of Health on behalf of the Northern Ireland Government and the Scottish Government. ASW and KBP are supported by the National Institute for Health Research Health Protection Research Unit (NIHR HPRU) in Healthcare Associated Infections and Antimicrobial Resistance at the University of Oxford in partnership with the UK Health Security Agency (UK HSA) (NIHR200915). ASW is also supported by the NIHR Oxford Biomedical Research Centre. KBP is also supported by the Huo Family Foundation. There are no other conflicts of interest.
Figures

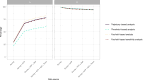
References
-
- World Health Organization. WHO Coronavirus (COVID-19) Dashboard. https://data.who.int/dashboards/covid19/cases (2024).
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

