An Innovative Binding-Protein-Based dsRNA Extraction Method: Comparison of Cost-Effectiveness of Virus Detection Methods Using High-Throughput Sequencing
- PMID: 40370068
- PMCID: PMC12415820
- DOI: 10.1111/1755-0998.14111
An Innovative Binding-Protein-Based dsRNA Extraction Method: Comparison of Cost-Effectiveness of Virus Detection Methods Using High-Throughput Sequencing
Abstract
Viral diseases represent a threat to global food production. Managing the impact of viruses on crop production requires the ability to monitor viruses, study their ecology and anticipate outbreaks. Double-stranded RNA (dsRNA) sequencing is a well-established and reliable method of detecting viruses and studying virome-host interactions and ecology. Compared to total RNA extraction, dsRNA extraction eliminates the majority of host RNAs, improving the recovery of viral RNAs. In this study, we developed and evaluated a novel dsRNA extraction method for high-throughput sequencing (HTS) applications based on the Flock House virus (FHV) B2 protein (B2-based method), and compared its performance with that of established cellulose-based and DRB4-based methods (commercial kit), as well as total RNA extraction techniques. The electrostatic properties of B2 have been instrumental in developing a bead-free and resin-free dsRNA extraction method. The B2-based method demonstrated high viral read recovery, achieving proportions exceeding 20% in most samples, and provided better dsRNA purity with less low weight molecule co-extracted RNA than the DRB4-based method and cellulose-based methods. Despite producing overall fewer total reads than the DRB4-based method, the B2-based enrichment for viral-derived dsRNA was better, with a higher percentage of viral reads, making it effective in virome profiling. Furthermore, it had an excellent detection specificity (0.97) and a good detection sensitivity (0.71), minimising false positives and false negatives. In addition, the B2-based method proved to be highly cost-effective, with a per-reaction cost of $4.47, compared to $35.34 for the DRB4-based method. This method offers a practical solution for laboratories with limited resources or for large-scale sampling for viral ecology studies. Future improvements to the B2-based method should focus on optimising sensitivity to Vitivirus species and developing scalable, automated workflows for high-throughput viral detection.
Keywords: HTS; Viromics; capture sequencing; dsRNA; grapevine virome; viral ecology.
© 2025 His Majesty the King in Right of Canada and The Author(s). Molecular Ecology Resources published by John Wiley & Sons Ltd. Reproduced with the permission of the Minister of Agriculture and Agri‐Food Canada.
Conflict of interest statement
Patent: The B2‐based dsRNA extraction method described in this manuscript is patented (WO2024086949A1). All the rights belong to the government of Canada.
The authors declare no conflicts of interest.
Figures
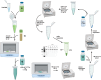

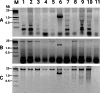
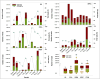
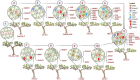



References
-
- Atsumi, G. , Sekine K. T., and Kobayashi K.. 2015. “A New Method to Isolate Total dsRNA.” Methods in Molecular Biology 1236: 27–37. - PubMed
-
- Cardwell, K. F. , Harmon C. L., Luster D. G., et al. 2023. “The Need and a Vision for a Diagnostic Assay Validation Network.” PhytoFrontiers 3: 9–17.
-
- Carisse, O. , Fall M. L., and Vincent C.. 2017. “Using a Biovigilance Approach for Pest and Disease Management in Quebec Vineyards.” Canadian Journal of Plant Pathology.
-
- Caruso, A. G. , Bertacca S., Ragona A., Matić S., Davino S., and Panno S.. 2022. “Evolutionary Analysis of Grapevine Virus A: Insights Into the Dispersion in Sicily (Italy).” Agriculture 12, no. 6: 835. 10.3390/agriculture12060835. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

