Bionectriaceae: a poorly known family of hypocrealean fungi with major commercial potential
- PMID: 40371418
- PMCID: PMC12070156
- DOI: 10.3114/sim.2025.111.04
Bionectriaceae: a poorly known family of hypocrealean fungi with major commercial potential
Abstract
The ascomycete family Bionectriaceae (Hypocreales) contains cosmopolitan species distributed throughout a broad range of environments, mainly occurring in terrestrial and freshwater ecosystems, with a less frequent occurrence in marine habitats. Members of the family are commonly used in industrial, pharmaceutical, and commercial applications. Applications utilise biodegraders and biocontrol agents, while certain taxa serve as a rich source of bioactive secondary metabolites. In recent years, several studies have proposed new taxonomic concepts within Bionectriaceae based on multi-gene phylogenetic inference. However, the status of several genera remains controversial or unclear, and many need to be re-collected and subjected to molecular analysis. The present study aims to improve our understanding of Bionectriaceae by re-examining CBS culture collection strains preliminarily identified as taxa within this family. Morphological and molecular phylogenetic analyses are based on alignments of the nuclear ribosomal subunits consisting of the internal transcribed spacer regions and intervening 5.8S nrDNA (ITS), as well as partial sequences for the 28S large subunit (LSU) nrDNA. Additional regions within protein-encoding genes were used, including the DNA-directed RNA polymerase II second largest subunit (RPB2), and translation elongation factor 1-alpha (TEF1) regions. The sequences generated were used to reconstruct a phylogenetic backbone of the family Bionectriaceae, and to delineate lineages and generic boundaries within it. Based on these results, seven new genera, 35 new species, and nine new combinations are proposed. A robustly supported phylogenetic framework is provided for Bionectriaceae, resolving 352 species and 50 well-supported genera. This study provides a solid foundation for more in-depth future studies on taxa in the family. Taxonomic novelties: New genera: Clavatomyces Lin Zhao & Crous, Collarimyces Lin Zhao & Crous, Vitreipilata Lin Zhao & Crous, Parageonectria Lin Zhao & Crous, Physaromyces Lin Zhao & Crous, Smyrniomyces Lin Zhao & Crous, Urticomyces Lin Zhao & Crous. New species: Acremonium paramultiramosum Lin Zhao & Crous, Clavatomyces prestoeae Lin Zhao & Crous, Clonostachys novocaledonica Lin Zhao & Crous, Clonostachys tropica Lin Zhao & Crous, Collarimyces guttiformis Lin Zhao & Crous, Emericellopsis mexicana Lin Zhao & Crous, Emericellopsis proliferata Lin Zhao & Crous, Emericellopsis soli Lin Zhao & Crous, Fusariella triangulispora Lin Zhao & Crous, Geonectria alni Lin Zhao & Crous, Geonectria quercus Lin Zhao & Crous, Geosmithia cupressina V. Meshram et al., Geosmithia magnispora Lin Zhao & Crous, Gliomastix olivacea Lin Zhao & Crous, Hydropisphaera armeniaca Lin Zhao & Crous, Hydropisphaera gossypina Lin Zhao & Crous, Hydropisphaera martinicensis Lin Zhao & Crous, Hydropisphaera solani Lin Zhao, L.W. Hou & Crous, Lasionectria chondroidea Lin Zhao & Crous, Lasionectria phormii Lin Zhao, L.W. Hou & Crous, Lasionectriopsis stereicola Lin Zhao & Crous, Nectriopsis cribrariae Lin Zhao & Crous, Nectriopsis floccosa Lin Zhao & Crous, Ovicillium theobromae Lin Zhao & Crous, Paracylindrocarpon jigongshanense Lin Zhao & Crous, Paracylindrocarpon spartinae Lin Zhao & Crous, Parageonectria arachispora Lin Zhao & Crous, Paragliomastix venezuelana Lin Zhao & Crous, Physaromyces sterilis Lin Zhao & Crous, Protocreopsis chlamydospora Lin Zhao & Crous, Protocreopsis gallica Lin Zhao & Crous, Roumegueriella echinulata Lin Zhao & Crous, Sesquicillium pouteriae Lin Zhao & Crous, Sesquicillium thailandense Lin Zhao & Crous, Smyrniomyces setaceus Lin Zhao & P.W. Crous. New combinations: Clavatomyces korfii (Lechat & J. Fourn.) Lin Zhao & Crous, Vitreipilata cirsii (Lechat & J. Fourn) Lin Zhao & Crous, Protocreopsis loweniae (Flakus et al.) Lin Zhao & Crous, Protocreopsis vulpina (Cooke) Lin Zhao & Crous, Proxiovicillium capsici (S.Q. Tong & Y.J. Wu) Lin Zhao & Crous, Sesquicillium shanghaiense (Zhang et al.) Lin Zhao & Crous, Urticomyces pseudoarenulus (Lechat & J. Fourn) Lin Zhao & Crous, Verruciconidia guizhouensis (S.Q. Tong & Y.J. Wu) Lin Zhao & Crous, Verruciconidia saulensis (Lechat & J. Fourn.) Lin Zhao & Crous. New synonyms: Clonostachys aquatica D.F. Bao et al., Emericellopsis microspora Backus & Orpurt. Citation: Zhao L, Groenewald JZ, Hou LW, Summerbell RC, Crous PW (2025). Bionectriaceae: a poorly known family of hypocrealean fungi with major commercial potential. Studies in Mycology 111: 115-198. doi: 10.3114/sim.2025.111.04.
Keywords: Bionectriaceae; industry; multi-locus phylogeny; new taxa; pharmaceutics; taxonomy.
© 2025 Westerdijk Fungal Biodiversity Institute.
Figures











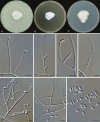
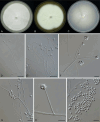


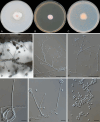
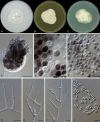


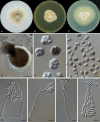
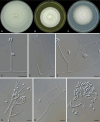
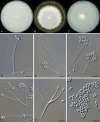
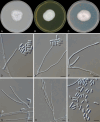
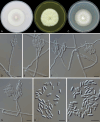


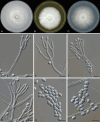


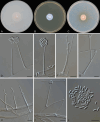







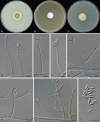




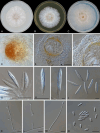

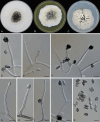
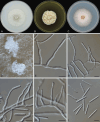

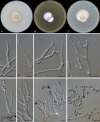
References
-
- Auer S, Ludwig-Müller J. (2014). Effects of the endophyte Acremonium alternatum on oilseed rape (Brassica napus) development and clubroot progression. Albanian Journal of Agricultural Sciences 13: 15–20.
-
- Auer S, Ludwig-Müller J. (2015). Biological control of clubroot (Plasmodiophora brassicae) by the endophytic fungus Acremonium alternatum. Journal of Endocytobiosis and Cell Research 26: 43–49.
LinkOut - more resources
Full Text Sources
