Deciphering the Molecular Crosstalk of Endoplasmic Reticulum Stress and Immune Infiltration in Endometriosis
- PMID: 40371728
- PMCID: PMC12079719
- DOI: 10.1111/aji.70092
Deciphering the Molecular Crosstalk of Endoplasmic Reticulum Stress and Immune Infiltration in Endometriosis
Abstract
Background: Endometriosis (EMs), characterized by ectopic endometrial growth causing infertility. Endoplasmic reticulum stress (ERS) is an important cellular defense mechanism. However, the correlation between ERS and EMs remains unclear. We aimed to investigate the relationship between them, identify biomarkers, and offer new insights into the treatment of EMs.
Methods: Two RNA expression datasets (GSE120103 and GSE25628) related to EMs in Homo sapiens were used to identify ERS-differentially expressed genes (ERS-DEGs). Protein-protein interaction (PPI) networks and CytoHubba (Cytoscape) identified ERS-associated HUB genes, with receiver operating characteristic curves (ROC) evaluating diagnostic value. Constructed mRNA-microRNA (miRNA)/RNA-transcription factor (TF) interaction networks and performed ssGSEA to compare immune infiltration between EMs patients and controls. Real-time quantitative polymerase chain reaction (RT-qPCR), western blotting (WB), and immunohistochemistry (IHC) were performed to assess potential biomarker levels.
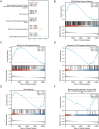
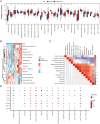
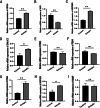
Results: Thirty-three ERS-DEGs were identified, with nine HUB genes (HSPA5, XBP1, HSP90B1, DNAJC3, PDIA3, PDIA6, PDIA4, HERPUD1, and MANF) demonstrating diagnostic efficacy (AUC > 0.7). Furthermore, immune infiltration revealed a significant relationship between immune cell abundance and HUB gene expression. Experimental validation confirmed the consistency of four biomarkers (HSPA5, HSP90B1, PDIA6, and HERPUD1). Regulatory network analysis identified 62 miRNAs and 44 TFs interacting with HUB genes, suggesting a multifactorial immunometabolic axis.
Conclusions: We identified four biomarkers (HSPA5, HSP90B1, PDIA6, and HERPUD1) associated with ERS that offer new insights into the detection and treatment of EMs. Our findings indicate an abnormal response to ERS and immune system infiltration contribute to the progression of EMs.
Keywords: endometriosis; endoplasmic reticulum stress; immune system infiltration.
© 2025 The Author(s). American Journal of Reproductive Immunology published by John Wiley & Sons Ltd.
Figures






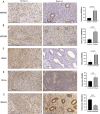
References
-
- Anglesio M. S. and Yong P. J., “Endometriosis‐Associated Ovarian Cancers,” Clinical Obstetrics and Gynecology 60, no. 4 (2017): 711–727. - PubMed
-
- Hetz C. and Papa F. R., “The Unfolded Protein Response and Cell Fate Control,” Molecular Cell 69, no. 2 (2018): 169–181. - PubMed
-
- Lin S., Yang L., Shi H., et al., “Endoplasmic Reticulum‐Targeting Photosensitizer Hypericin Confers Chemo‐Sensitization Towards Oxaliplatin Through Inducing Pro‐Death Autophagy,” International Journal of Biochemistry & Cell Biology 87 (2017): 54–68. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

