Exploring the Relationship of Transposable Elements and Ageing: Causes and Consequences
- PMID: 40373205
- PMCID: PMC12127037
- DOI: 10.1093/gbe/evaf088
Exploring the Relationship of Transposable Elements and Ageing: Causes and Consequences
Abstract
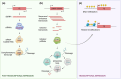
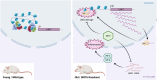
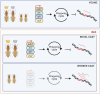
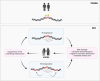
Ageing is a gradual biological process marked by a decline in physiological function, increasing susceptibility to disease, and mortality. Transposable elements (TEs) are repetitive DNA sequences capable of moving within the genome and thus potentially inducing mutations and disrupting normal cellular functions. Their mobile nature contributes to genomic variation, as transposition events can alter gene expression, chromosome structure, and the epigenetic landscape. To mitigate TE-induced damage, cells rely on epigenetic mechanisms, such as DNA methylation, histone modifications, and small RNAs, to repress TE activity. However, these silencing mechanisms become less effective with age, leading to increased TE activation. This review explores the dual role of TEs as both a cause and consequence of ageing, suggesting a complex relationship between TEs and the ageing process.
Keywords: age-related diseases; chromatin; long-lived organisms; senescence; transposable elements.
© The Author(s) 2025. Published by Oxford University Press on behalf of Society for Molecular Biology and Evolution.
Figures

References
-
- Bendall ML, de Mulder M, Iñiguez LP, Lecanda-Sánchez A, Pérez-Losada M, Ostrowski MA, Jones RB, Mulder LCF, Reyes-Terán G, Crandall KA, et al. Telescope: characterization of the retrotranscriptome by accurate estimation of transposable element expression. PLoS Comput Biol. 2019:15(9):e1006453. 10.1371/journal.pcbi.1006453. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

