Comparative transcriptome analysis reveals key genes associated with meiotic stability and high seed setting rate in tetraploid rice
- PMID: 40375090
- PMCID: PMC12080012
- DOI: 10.1186/s12870-025-06672-x
Comparative transcriptome analysis reveals key genes associated with meiotic stability and high seed setting rate in tetraploid rice
Abstract
Background: Polyploid rice has a high yield potential and excellent nutritional quality. The development of polyploid rice remained critically limited for several decades due to low seed setting rate until the successful breeding of polyploid meiosis stability (PMeS) lines. To determine the mechanism responsible for meiotic stability and high seed setting rate of PMeS line, agronomic traits, pollen fertility and viability, and meiotic behaviors of PMeS and non-PMeS lines were investigated. Further, comparative transcriptome analysis was performed to identify genes associated with meiotic stability and high seed setting rate in PMeS line.
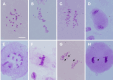
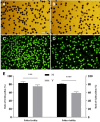
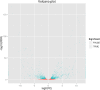
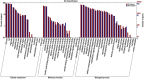
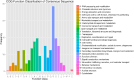
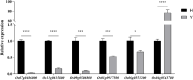
Results: The seed setting rate, fertile and viable pollen ratios of PMeS line were significantly higher than those of non-PMeS line. The PMeS line exhibited stable meiosis, and chromosomes mainly paired as bivalents, rarely as univalents and multivalents in prophase I. Few lagging chromosomes were observed in anaphase I. By contrast, the homologous chromosomes pairing was disorganized in the non-PMeS line, with low frequencies of bivalents and high frequencies of univalents and multivalents in prophase I, while more cells with increased lagging chromosomes were detected in anaphase I. Many differentially expressed genes (DEGs) between PMeS and non-PMeS lines were identified through comparative transcriptome analysis. Some meiosis-related genes were specifically investigated from all DEGs. Further, several meiotic genes were identified as candidate genes.
Conclusions: The study not only demonstrates the morphological, cytological, and molecular differences between the PMeS and non-PMeS lines, but also provides several key genes associated with meiotic stability and high seed setting rate in tetraploid rice.
Keywords: High seed setting rate; Meiotic genes; Meiotic stability; Tetraploid rice; Transcriptome analysis.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
The breeding of two polyploid rice lines with the characteristic of polyploid meiosis stability.Sci China C Life Sci. 2007 Jun;50(3):356-66. doi: 10.1007/s11427-007-0049-6. Sci China C Life Sci. 2007. PMID: 17609893
-
Cytological observation and transcriptome analysis reveal that NTFR1 is a new tetraploid rice fertility gene using the tetraploid fertility-directed lines.Plant Sci. 2025 Jun;355:112437. doi: 10.1016/j.plantsci.2025.112437. Epub 2025 Feb 28. Plant Sci. 2025. PMID: 40024612
-
Cytological Observations and Bulked-Segregant Analysis Coupled Global Genome Sequencing Reveal Two Genes Associated with Pollen Fertility in Tetraploid Rice.Int J Mol Sci. 2021 Jan 15;22(2):841. doi: 10.3390/ijms22020841. Int J Mol Sci. 2021. PMID: 33467721 Free PMC article.
-
Fertile Tetraploids: New Resources for Future Rice Breeding?Front Plant Sci. 2020 Aug 11;11:1231. doi: 10.3389/fpls.2020.01231. eCollection 2020. Front Plant Sci. 2020. PMID: 32849760 Free PMC article. Review.
-
A New Way of Rice Breeding: Polyploid Rice Breeding.Plants (Basel). 2021 Feb 24;10(3):422. doi: 10.3390/plants10030422. Plants (Basel). 2021. PMID: 33668223 Free PMC article. Review.
References
-
- Comai L. The advantages and disadvantages of being polyploid. Nat Rev Genet. 2005;6:836–46. - PubMed
-
- Soltis DE, Albert VA, Leebens-Mack J, Bell CD, Paterson AH, Zheng C, et al. Polyploidy and angiosperm diversification. Am J Bot. 2009;96:336–48. - PubMed
-
- Fang Z, Morrell PL. Domestication: polyploidy boosts domestication. Nat Plants. 2016;2:16116. - PubMed
-
- Yu H, Lin T, Meng X, Du H, Zhang J, Liu G, et al. A route to de Novo domestication of wild allotetraploid rice. Cell. 2021;184:1156–70. - PubMed
-
- Otto SP. The evolutionary consequences of polyploidy. Cell. 2007;131:452–62. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

