Machine learning-based multimodal radiomics and transcriptomics models for predicting radiotherapy sensitivity and prognosis in esophageal cancer
- PMID: 40381695
- PMCID: PMC12221354
- DOI: 10.1016/j.jbc.2025.110242
Machine learning-based multimodal radiomics and transcriptomics models for predicting radiotherapy sensitivity and prognosis in esophageal cancer
Abstract
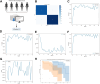
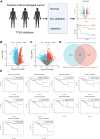
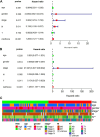
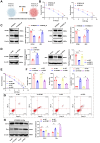
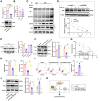
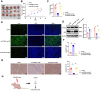
Radiotherapy plays a critical role in treating esophageal cancer, but individual responses vary significantly, impacting patient outcomes. This study integrates machine learning-driven multimodal radiomics and transcriptomics to develop predictive models for radiotherapy sensitivity and prognosis in esophageal cancer. We applied the SEResNet101 deep learning model to imaging and transcriptomic data from the UCSC Xena and TCGA databases, identifying prognosis-associated genes such as STUB1, PEX12, and HEXIM2. Using Lasso regression and Cox analysis, we constructed a prognostic risk model that accurately stratifies patients based on survival probability. Notably, STUB1, an E3 ubiquitin ligase, enhances radiotherapy sensitivity by promoting the ubiquitination and degradation of SRC, a key oncogenic protein. In vitro and in vivo experiments confirmed that STUB1 overexpression or SRC silencing significantly improves radiotherapy response in esophageal cancer models. These findings highlight the predictive power of multimodal data integration for individualized radiotherapy planning and underscore STUB1 as a promising therapeutic target for enhancing radiotherapy efficacy in esophageal cancer.
Keywords: SEResNet101; SRC ubiquitination; STUB1; esophageal cancer; prognostic risk model; radiotherapy sensitivity.
Copyright © 2025 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that there are no conflicts of interests with the contents of this article.
Figures


Similar articles
-
Characterization of novel anoikis-related genes as prognostic biomarkers and key determinants of the immune microenvironment in esophageal cancer.Front Immunol. 2025 Jul 11;16:1599171. doi: 10.3389/fimmu.2025.1599171. eCollection 2025. Front Immunol. 2025. PMID: 40746552 Free PMC article.
-
Comparison of Two Modern Survival Prediction Tools, SORG-MLA and METSSS, in Patients With Symptomatic Long-bone Metastases Who Underwent Local Treatment With Surgery Followed by Radiotherapy and With Radiotherapy Alone.Clin Orthop Relat Res. 2024 Dec 1;482(12):2193-2208. doi: 10.1097/CORR.0000000000003185. Epub 2024 Jul 23. Clin Orthop Relat Res. 2024. PMID: 39051924
-
Computed tomography-based radiomics predicts prognostic and treatment-related levels of immune infiltration in the immune microenvironment of clear cell renal cell carcinoma.BMC Med Imaging. 2025 Jul 1;25(1):213. doi: 10.1186/s12880-025-01749-3. BMC Med Imaging. 2025. PMID: 40596969 Free PMC article.
-
MRI-Based Radiomics Methods for Predicting Ki-67 Expression in Breast Cancer: A Systematic Review and Meta-analysis.Acad Radiol. 2024 Mar;31(3):763-787. doi: 10.1016/j.acra.2023.10.010. Epub 2023 Nov 2. Acad Radiol. 2024. PMID: 37925343
-
Cost-effectiveness of using prognostic information to select women with breast cancer for adjuvant systemic therapy.Health Technol Assess. 2006 Sep;10(34):iii-iv, ix-xi, 1-204. doi: 10.3310/hta10340. Health Technol Assess. 2006. PMID: 16959170
References
-
- DiSiena M., Perelman A., Birk J., Rezaizadeh H. Esophageal cancer: an updated review. South. Med. J. 2021;114:161–168. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

