A sexual/apomictic consensus linkage map of Eragrostis curvula at tetraploid level
- PMID: 40382602
- PMCID: PMC12085070
- DOI: 10.1186/s12870-025-06676-7
A sexual/apomictic consensus linkage map of Eragrostis curvula at tetraploid level
Abstract
Background: Apomixis is an asexual reproduction process that allows plants to bypass meiosis and fertilization, resulting in clonal seeds that are genetically identical to the maternal genotype. Eragrostis curvula is a grass species used as model to disclose the mechanism associated to diplosporous apomixis. Previously, the first E. curvula linkage maps were developed using a F1 population derived from a cross between a sexual female parent (cv. OTA-S) and a facultative apomictic pollen donor (cv. Don Walter). Even though this work allows the identification a markers linked to apomixis in the male parent, the number of hybrids was not enough to produce a consensus map. Here, a new population is presented, increasing the number of genotyped hybrids to 107 which allows the construction of a consensus map and the development of KASP markers.
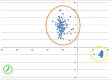
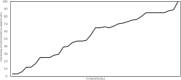
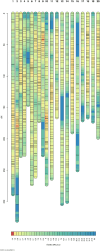
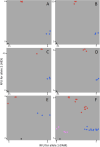
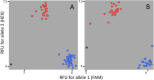
Results: We constructed a consensus linkage map at the tetraploid level using a mapping population segregating for reproductive mode. Within this map, a region associated with apomeiosis (the APO locus) was identified using maternal and paternal SNP markers, along with three paternal markers that exhibited strong linkage with the trait. KASP markers were developed, one of which demonstrated 100% concordance with the cytoembryological phenotype of individuals in both the mapping population and other E. curvula genotypes. Through synteny analysis, the APO locus was mapped onto the E. curvula reference genome, and two genes that could be part of molecular pathways involved in apomeiosis were proposed.
Conclusions: This study presents the first consensus genetic map and the development of KASP markers for phenotyping reproductive modes in E. curvula. This map enables the association of the apomixis-determining region with molecular markers from both parental genotypes, including the reference sexual tetraploid genotype of the species (OTA-S). The development and validation of co-dominant molecular KASP markers linked to the APO locus provide a crucial tool for future research and breeding.
Keywords: Apomeiosis; Apomixis; Mapping; Weeping lovegrass.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures


References
-
- Akiyama Y, Hanna W, Ozias-Akins P. High-resolution physical mapping reveals that the apospory-specific genomic region (ASGR) in Cenchrus ciliaris is located on a heterochromatic and hemizygous region of a single chromosome. Theor Appl Genet. 2005;111:1042–51. 10.1007/s00122-005-0020-5. - DOI - PubMed
-
- Albertini E, Barcaccia G, Mazzucato A, Sharbel TF, Falcinelli M. Apomixis in the era of biotechnology. In: Pua E, Davey M, editors. Plant Developmental Biology - Biotechnological Perspectives. Berlin: Springer; 2010. p. 405–36. 10.1007/978-3-642-02301-9_20.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

