Unlocking complexity through neutron scattering: Structure and dynamics of protein-polymer conjugates
- PMID: 40384628
- PMCID: PMC12086510
- DOI: 10.1002/pro.70137
Unlocking complexity through neutron scattering: Structure and dynamics of protein-polymer conjugates
Abstract
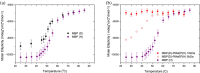
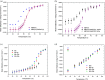
Protein-polymer conjugates are engineered to enhance the pharmacokinetics and stability of biopharmaceuticals. This review delves into the structural and dynamic characteristics of protein-polymer conjugates, employing advanced techniques such as neutron scattering, circular dichroism, and fluorescence spectroscopy. Here, we focus on model proteins like maltose-binding protein (MBP), bovine serum albumin (BSA), and myoglobin (Mb) conjugated with hydrolysable polyphosphoesters (PPEs) to generate fully degradable polymer-protein conjugates. The study underscores the influence of factors such as the polymers' molar mass, grafting density, hydration levels, and deuteration on protein stability, flexibility, and thermal properties. The study demonstrates that the degree of polymerization, solvation properties, and isotopic composition play crucial roles in determining the behavior of protein-polymer complexes. In particular, neutron scattering techniques are invaluable in unraveling the interaction mechanisms, providing valuable insights that can inform the optimization of protein-polymer conjugates for therapeutic applications.
Keywords: dynamical fluctuation; isotopic deuteration; neutron scattering; protein–polymer conjugates; radius of gyration.
© 2025 The Author(s). Protein Science published by Wiley Periodicals LLC on behalf of The Protein Society.
Figures
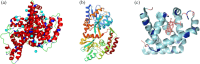
Similar articles
-
Investigation of the structure of protein-polymer conjugates in solution reveals the impact of protein deuteration and the size of the polymer on its thermal stability.Protein Sci. 2024 Jun;33(6):e5032. doi: 10.1002/pro.5032. Protein Sci. 2024. PMID: 38801224 Free PMC article.
-
Conformation of Myoglobin-Poly(Ethyl Ethylene Phosphate) Conjugates Probed by SANS: Correlation with Polymer Grafting Density and Interaction.Macromol Biosci. 2021 Feb;21(2):e2000356. doi: 10.1002/mabi.202000356. Epub 2021 Jan 4. Macromol Biosci. 2021. PMID: 33393176
-
Investigation into the Relaxation Dynamics of Polymer-Protein Conjugates Reveals Surprising Role of Polymer Solvation on Inherent Protein Flexibility.Biomacromolecules. 2016 Jan 11;17(1):141-7. doi: 10.1021/acs.biomac.5b01269. Epub 2015 Dec 4. Biomacromolecules. 2016. PMID: 26568153
-
Neutron Scattering from Polymers: Five Decades of Developing Possibilities.Annu Rev Chem Biomol Eng. 2016 Jun 7;7:1-28. doi: 10.1146/annurev-chembioeng-080615-034429. Annu Rev Chem Biomol Eng. 2016. PMID: 27276548 Review.
-
Structural Investigations of Protein-Lipid Complexes Using Neutron Scattering.Methods Mol Biol. 2019;2003:201-251. doi: 10.1007/978-1-4939-9512-7_11. Methods Mol Biol. 2019. PMID: 31218621 Review.
References
-
- Butler P, Doucet M, Jackson A, King S. SasView. 2012. Available from: http://www.sasview.org/
-
- DeWitt SH, Maryanoff BE. Deuterated drug molecules: focus on FDA‐approved Deutetrabenazine. Biochemistry. 2018;57:472–473. - PubMed
-
- Doster W, Cusack S, Petry W. Dynamical transition of myoglobin revealed by inelastic neutron scattering. Nature. 1989;337:754–756. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

