Surveying Enzyme Crystal Structures Reveals the Commonality of Active-Site Solvent Accessibility and Enzymatic Water Networks
- PMID: 40385134
- PMCID: PMC12079200
- DOI: 10.1021/acsomega.4c10721
Surveying Enzyme Crystal Structures Reveals the Commonality of Active-Site Solvent Accessibility and Enzymatic Water Networks
Abstract
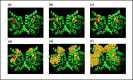
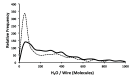
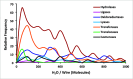
Despite the demonstrable dependence of enzyme functionality on solvation, the notion of water being directly chemically required for catalysis inside active sites remains unexplored. Here we report that over 99% of 1013 enzyme crystals obtained by X-ray crystallography with high resolution (<1.5 Å) contain continuous chains of water linking residues within the active site to bulk water. Also reported are the findings which inspired this study-that electric fields experienced by water hydrogen atoms are on average twice as strong in the active sites of both chains of bacterial polynucleotide kinase (PDB 4QM6) structures compared to those in bulk water. These results point to the possibility that water molecules within active sites may be paramount to the immense catalytic power of enzymes, especially for mechanisms requiring hydronium or hydroxide ions.
© 2025 The Authors. Published by American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures




Similar articles
-
"Seeing Is Believing": How Neutron Crystallography Informs Enzyme Mechanisms by Visualizing Unique Water Species.Biology (Basel). 2024 Oct 22;13(11):850. doi: 10.3390/biology13110850. Biology (Basel). 2024. PMID: 39596805 Free PMC article. Review.
-
NMR crystallography of enzyme active sites: probing chemically detailed, three-dimensional structure in tryptophan synthase.Acc Chem Res. 2013 Sep 17;46(9):2008-17. doi: 10.1021/ar3003333. Epub 2013 Mar 28. Acc Chem Res. 2013. PMID: 23537227 Free PMC article.
-
Comparison of x-ray crystal structures of an acyl-enzyme intermediate of subtilisin Carlsberg formed in anhydrous acetonitrile and in water.Proc Natl Acad Sci U S A. 1998 Oct 27;95(22):12918-23. doi: 10.1073/pnas.95.22.12918. Proc Natl Acad Sci U S A. 1998. PMID: 9789015 Free PMC article.
-
Ionized water confined in graphene nanochannels.Phys Chem Chem Phys. 2019 May 8;21(18):9285-9295. doi: 10.1039/c9cp00075e. Phys Chem Chem Phys. 2019. PMID: 30931451
-
Orotidine 5'-Monophosphate Decarboxylase: Probing the Limits of the Possible for Enzyme Catalysis.Acc Chem Res. 2018 Apr 17;51(4):960-969. doi: 10.1021/acs.accounts.8b00059. Epub 2018 Mar 29. Acc Chem Res. 2018. PMID: 29595949 Free PMC article. Review.
References
-
- Agmon N. The grotthuss mechanism. Chem. Phys. Lett. 1995, 244 (5–6), 456–462. 10.1016/0009-2614(95)00905-J. - DOI
LinkOut - more resources
Full Text Sources
