Investigating the impact of synonymous gene recoding on a recombinantly expressed monoclonal antibody under different process parameters
- PMID: 40385532
- PMCID: PMC12079346
- DOI: 10.1002/btm2.10750
Investigating the impact of synonymous gene recoding on a recombinantly expressed monoclonal antibody under different process parameters
Abstract
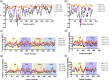
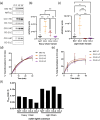
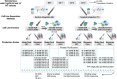
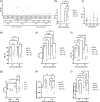
Monoclonal antibodies (mAbs) are commonly used biologic therapeutics with a wide variety of clinical applications. During the development process, manufacturers consider different production parameters to improve protein yield and achieve appropriate quality of the product. Synonymous gene recoding is one of such attributes that is often considered and implemented to enhance protein expression. However, it has to be used with caution, as it may lead to protein misfolding and ER stress, which complicates efforts to manufacture the desired mAb. To investigate how changing mRNA sequence composition under different protein production parameters might affect the quality of recombinantly produced mAbs, we performed a comprehensive and systematic study assessing impact of synonymous gene recoding (commonly referred to as codon optimization) strategies in the context of varied cell culture parameters on product quality, biochemical and functional characteristics. We report the impact of these parameters on mAb glycosylation profiles, charge variant profile, aggregation, fragmentation, and mAb functional response from combinations of different production parameters. These results uncovered a complex interplay of sequence composition and manufacturing parameters and emphasize the importance of assessing changes to key quality attributes when optimizing mAb manufacturing, including the use of synonymous gene recoding.
Keywords: CHO cells; codon optimization; monoclonal antibody; process changes.
© 2025 The Author(s). Bioengineering & Translational Medicine published by Wiley Periodicals LLC on behalf of The American Institute of Chemical Engineers.
Conflict of interest statement
Anton A. Komar is a founder of SATOR Therapeutics LLC. Anton A. Komar's spouse is a founder of DAPCEL, Inc. and SATOR Therapeutics LLC. The other authors declare no competing interests.
Figures

Similar articles
-
Short-Term Memory Impairment.2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Jun 8. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 31424720 Free Books & Documents.
-
A rapid and systematic review of the clinical effectiveness and cost-effectiveness of paclitaxel, docetaxel, gemcitabine and vinorelbine in non-small-cell lung cancer.Health Technol Assess. 2001;5(32):1-195. doi: 10.3310/hta5320. Health Technol Assess. 2001. PMID: 12065068
-
Management of urinary stones by experts in stone disease (ESD 2025).Arch Ital Urol Androl. 2025 Jun 30;97(2):14085. doi: 10.4081/aiua.2025.14085. Epub 2025 Jun 30. Arch Ital Urol Androl. 2025. PMID: 40583613 Review.
-
The Black Book of Psychotropic Dosing and Monitoring.Psychopharmacol Bull. 2024 Jul 8;54(3):8-59. Psychopharmacol Bull. 2024. PMID: 38993656 Free PMC article. Review.
-
Sexual Harassment and Prevention Training.2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. 2024 Mar 29. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025 Jan–. PMID: 36508513 Free Books & Documents.
References
-
- Urquhart L. Top companies and drugs by sales in 2022. Nat Rev Drug Discov. 2023;22(4):260. - PubMed
-
- Hodgson J. Drug pipeline 3Q23 – ERT, bispecifics and CRISPR in sickle cell disease. Nat Biotechnol. 2023;41(11):1498‐1500. - PubMed
-
- Jarasch A, Koll H, Regula JT, Bader M, Papadimitriou A, Kettenberger H. Developability assessment during the selection of novel therapeutic antibodies. J Pharm Sci. 2015;104(6):1885‐1898. - PubMed
-
- Li Z, Du X, Wang YC. A survey of FDA approved monoclonal antibodies and fc‐fusion proteins for manufacturing changes and comparability assessment. Pharm Res. 2024;41(1):13‐27. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

