Integrated ATAC-seq and RNA-seq analysis identifies key regulatory elements in NK cells activated with feeder cells and IL-2
- PMID: 40385536
- PMCID: PMC12079432
- DOI: 10.1002/btm2.10747
Integrated ATAC-seq and RNA-seq analysis identifies key regulatory elements in NK cells activated with feeder cells and IL-2
Abstract
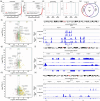
Natural killer (NK) cells are in development for allogeneic immunotherapy; however, for such use as off-the-shelf medicines, NK cells need to undergo ex vivo expansion, typically through activation with feeder cells and cytokines, to generate sufficient cells for clinical applications. Upon stimulation with feeder cells in the presence of cytokines, NK cells undergo profound changes in gene expression, altering their metabolic activity, cell cycle progression, and growth behavior, but the precise changes that drive this transformation remain poorly understood. In this study, we identified significant differences in the transcriptome and chromatin accessibility of NK cells 7 days after feeder cell and cytokine activation, with the changes even more pronounced in genome regions closer to enhancers. Several transcription factors, including AP-1, IRF4, STATs, T-bet, Eomes, and bHLHE40, which play key roles in NK cell development and immune response, exhibited differential binding activity between unstimulated and day 7 NK cells. Gene sets composed of target genes downstream of these transcription factors were also enriched at day 7, implying their involvement in NK cell activation. Moreover, we compared potential super-enhancer regions in NK cells before and after activation, combined with the transcriptional activity of nearby genes. We identified stable and transcriptionally active super-enhancers in unstimulated and day 7 NK cells, as well as those that form or disappear after co-culture initiation. The transcriptomic and epigenetic characterization of NK cells presented in this study could facilitate the ex vivo expansion and engineering of functionally superior NK cells.
Keywords: ATAC‐seq; NK cells; feeder cells; super‐enhancers; transcription factor.
© 2025 The Author(s). Bioengineering & Translational Medicine published by Wiley Periodicals LLC on behalf of American Institute of Chemical Engineers.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Optimization of Human NK Cell Manufacturing: Fully Automated Separation, Improved Ex Vivo Expansion Using IL-21 with Autologous Feeder Cells, and Generation of Anti-CD123-CAR-Expressing Effector Cells.Hum Gene Ther. 2017 Oct;28(10):897-913. doi: 10.1089/hum.2017.157. Epub 2017 Aug 15. Hum Gene Ther. 2017. PMID: 28810809
-
Superior Expansion and Cytotoxicity of Human Primary NK and CAR-NK Cells from Various Sources via Enriched Metabolic Pathways.Mol Ther Methods Clin Dev. 2020 Jun 24;18:428-445. doi: 10.1016/j.omtm.2020.06.014. eCollection 2020 Sep 11. Mol Ther Methods Clin Dev. 2020. PMID: 32695845 Free PMC article.
-
Feeder-cell-free system for ex vivo production of natural killer cells from cord blood hematopoietic stem and progenitor cells.Front Immunol. 2025 Feb 20;16:1531736. doi: 10.3389/fimmu.2025.1531736. eCollection 2025. Front Immunol. 2025. PMID: 40051631 Free PMC article.
-
Shaping of Natural Killer Cell Antitumor Activity by Ex Vivo Cultivation.Front Immunol. 2017 Apr 26;8:458. doi: 10.3389/fimmu.2017.00458. eCollection 2017. Front Immunol. 2017. PMID: 28491060 Free PMC article. Review.
-
Building a Better Defense: Expanding and Improving Natural Killer Cells for Adoptive Cell Therapy.Cells. 2024 Mar 5;13(5):451. doi: 10.3390/cells13050451. Cells. 2024. PMID: 38474415 Free PMC article. Review.
References
-
- Prager I, Watzl C. Mechanisms of natural killer cell‐mediated cellular cytotoxicity. J Leukoc Biol. 2019;105(6):1319‐1329. - PubMed
LinkOut - more resources
Full Text Sources

