PRDX1 protects ATM from arsenite-induced proteotoxicity and maintains its stability during DNA damage signaling
- PMID: 40387816
- PMCID: PMC12088036
- DOI: 10.18632/oncotarget.28720
PRDX1 protects ATM from arsenite-induced proteotoxicity and maintains its stability during DNA damage signaling
Abstract
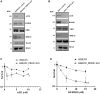
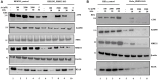
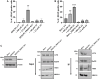
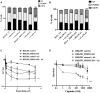
Redox regulation and DNA repair coordination are essential for genomic stability. Peroxiredoxin 1 (PRDX1) is a thiol-dependent peroxidase and a chaperone that protects proteins from excessive oxidation. ATM kinase (Ataxia-Telangiectasia Mutated) and the MRN (MRE11-RAD50-NBS1) complex are DNA damage signaling and repair proteins. We previously showed that cells lacking PRDX1 are sensitive to arsenite, a toxic metal that induces DNA single- and double-strand breaks (DSBs). Herein, we showed that PRDX1 interacts with ATM. PRDX1-deleted cells have reduced ATM, MRE11, and RAD50 protein levels, but not NBS1. In control cells treated with arsenite, we observed γH2AX foci formation due to arsenite-induced DSBs, and not from PRDX1-deleted cells. Arsenite caused profound depletion of ATM in PRDX1-deleted cells, suggesting that PRDX1 protects and stabilizes ATM required to form γH2AX foci. Importantly, arsenite pretreatment of PRDX1-deleted cells caused hypersensitivity to chemotherapeutic agents that generate DSBs. Analysis of a clinical cohort of ovarian cancers treated with platinum chemotherapy revealed that tumours with high PRDX1/high ATM or high PRDX1/high MRE11 expression manifested aggressive phenotypes and poor patient survival. The data suggest that PRDX1 can predict responses to chemotherapy, and targeting PRDX1 could be a viable strategy to improve the efficacy of platinum chemotherapy.
Keywords: cell cycle; homologous recombination; protein interaction; protein modification; redox signaling.
Conflict of interest statement
Authors have no conflicts of interest to declare.
Figures



Similar articles
-
Loss of the tumor suppressor BIN1 enables ATM Ser/Thr kinase activation by the nuclear protein E2F1 and renders cancer cells resistant to cisplatin.J Biol Chem. 2019 Apr 5;294(14):5700-5719. doi: 10.1074/jbc.RA118.005699. Epub 2019 Feb 7. J Biol Chem. 2019. PMID: 30733337 Free PMC article.
-
Rad17 recruits the MRE11-RAD50-NBS1 complex to regulate the cellular response to DNA double-strand breaks.EMBO J. 2014 Apr 16;33(8):862-77. doi: 10.1002/embj.201386064. Epub 2014 Feb 16. EMBO J. 2014. PMID: 24534091 Free PMC article.
-
MRE11 UFMylation promotes ATM activation.Nucleic Acids Res. 2019 May 7;47(8):4124-4135. doi: 10.1093/nar/gkz110. Nucleic Acids Res. 2019. PMID: 30783677 Free PMC article.
-
ATP puts the brake on DNA double-strand break repair: a new study shows that ATP switches the Mre11-Rad50-Nbs1 repair factor between signaling and processing of DNA ends.Bioessays. 2014 Dec;36(12):1170-8. doi: 10.1002/bies.201400102. Epub 2014 Sep 11. Bioessays. 2014. PMID: 25213441 Review.
-
ATM-Dependent Phosphorylation of All Three Members of the MRN Complex: From Sensor to Adaptor.Biomolecules. 2015 Oct 23;5(4):2877-902. doi: 10.3390/biom5042877. Biomolecules. 2015. PMID: 26512707 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

