B cell dysfunction in thalamus and brainstem involvement and high lactate caused by novel mutation of EARS2 gene
- PMID: 40389993
- PMCID: PMC12090667
- DOI: 10.1186/s13052-025-01999-5
B cell dysfunction in thalamus and brainstem involvement and high lactate caused by novel mutation of EARS2 gene
Abstract
Purpose: The EARS2 gene, a member of the mt-aaRS family, encodes mitochondrial glutamyl-tRNA synthetase (GluRS), which is involved in the synthesis of mitochondrial proteins. Pathogenic defects in EARS2 may cause mitochondrial OXPHOS deficiency, which is associated with a rare autosomal-recessive mitochondrial disease, leukoencephalopathy with thalamus and brainstem involvement and high lactate (LTBL).
Methods: In this study, clinical features were obtained, and whole-exome sequencing was conducted on a patient with LTBL. B- and T-cell immunophenotyping and protein expression were analyzed using flow cytometry, and B-cell metabolism was investigated using confocal microscopy.
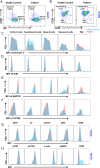
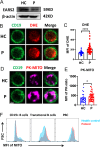
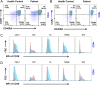
Results: The patient with LTBL exhibited typical neurological manifestations, recurrent respiratory tract infections, and humoral immune disorders. Molecular analysis revealed a compound heterozygous novel mutation in c.1304T > A (p.L435Q) and a previously reported c.319 C > T (p.R107C) mutation of EARS2. The mutations led to protein structural modifications of EARS2. The patient also exhibited disrupted peripheral B-cell differentiation and B-cell receptor signal transduction. The EARS2 mutation led to decreased expression of CD38 and dysfunction of mitochondrial metabolism, with elevated reactive oxygen species levels in B cells.
Conclusion: We identified a novel mutation of the EARS2 gene in a patient with LTBL, expanding the mutation database. The mutation of EARS2 modified protein structure and impaired B-cell function, decreased CD38 expression, and led to dysfunction of mitochondrial metabolism, all of which may account for the recurrent respiratory tract infections and humoral immune disorders observed in LTBL.
Keywords: B cell; BCR signal; EARS2; Gene mutation; LTBL.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The study was authorized by the Ethics Committee of Tongji Hospital, Tongji Medical College, Huazhong University of Science and Technology, Wuhan, China (TJ-IRB202407105). Consent to participate: Parents of the patient provided written informed consent to participation. Conflicts of interest: The authors have no relevant financial or non-financial interests to disclose.
Figures


Similar articles
-
Identification of a Novel Variant in EARS2 Associated with a Severe Clinical Phenotype Expands the Clinical Spectrum of LTBL.Genes (Basel). 2020 Sep 2;11(9):1028. doi: 10.3390/genes11091028. Genes (Basel). 2020. PMID: 32887222 Free PMC article.
-
A compound heterozygous EARS2 mutation associated with mild leukoencephalopathy with thalamus and brainstem involvement and high lactate (LTBL).Brain Dev. 2016 Oct;38(9):857-61. doi: 10.1016/j.braindev.2016.04.002. Epub 2016 Apr 23. Brain Dev. 2016. PMID: 27117034
-
Leukoencephalopathy with thalamus and brainstem involvement and high lactate caused by novel mutations in the EARS2 gene in two siblings.J Neurol Sci. 2016 Jun 15;365:54-8. doi: 10.1016/j.jns.2016.04.008. Epub 2016 Apr 9. J Neurol Sci. 2016. PMID: 27206875
-
Mitochondrial aminoacyl-tRNA synthetase disorders: an emerging group of developmental disorders of myelination.J Neurodev Disord. 2019 Dec 16;11(1):29. doi: 10.1186/s11689-019-9292-y. J Neurodev Disord. 2019. PMID: 31839000 Free PMC article. Review.
-
Mitochondrial syndromes with leukoencephalopathies.Semin Neurol. 2012 Feb;32(1):55-61. doi: 10.1055/s-0032-1306387. Epub 2012 Mar 15. Semin Neurol. 2012. PMID: 22422207 Review.
References
-
- Chan DC. Mitochondrial dynamics and its involvement in disease. Annu Rev Pathol. 2020;15:235–59. - PubMed
-
- Sissler M, González-Serrano LE, Westhof E. Recent advances in mitochondrial Aminoacyl-tRNA synthetases and disease. Trends Mol Med. 2017;23:693–708. - PubMed
-
- Antonellis A, Green ED. The role of aminoacyl-tRNA synthetases in genetic diseases. Annu Rev Genomics Hum Genet. 2008;9:87–107. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

