Self and parasite-derived peptides selected upon DERAA-bearing HLA-DRB1 alleles activate CD4+ T cells from Chagas cardiomyopathy patients and are associated with ventricular dysfunction
- PMID: 40391216
- PMCID: PMC12086171
- DOI: 10.3389/fimmu.2025.1527115
Self and parasite-derived peptides selected upon DERAA-bearing HLA-DRB1 alleles activate CD4+ T cells from Chagas cardiomyopathy patients and are associated with ventricular dysfunction
Abstract
Introduction: Human infection with the protozoan Trypanosoma cruzi causes Chagas disease, which may lead to a deadly dilated cardiomyopathy resulting from T-cell-mediated inflammation. We found that specific HLA-DRB1 alleles (*0103, *0402, *1301, and *1302) that display the DERAA motif are linked to this severe clinical manifestation of Chagas disease.
Methods: We employed computational analysis, in vitro functional assays, and single-cell RNA sequencing to determine the response of CD4+ T cells from indeterminate (IND) and cardiac (CCC) Chagas patients to peptides selected on DERAA-bearing alleles.
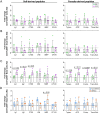
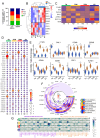
Results: We observed that these alleles display binding affinity towards host-derived peptides with sequence similarity to parasite-derived proteins. These peptides can activate and induce proliferation of CD4+ T-cells from CCC, but not IND. Importantly, the magnitude of this response correlated with the severity of ventricular dysfunction and increased production of soluble factors associated with myocardial fibrosis. Analysis of differentially expressed genes (DEGs) in activated CD4+ T-cells from individuals with the DERAA-DRB1 alleles demonstrated a high expression of cytotoxic, chemotactic and proapoptotic genes, linking these cells with pathogenic functions. Finally, we observed the upregulation of genes that code for the host proteins that contain the potentially pathogenic peptides in the cardiac tissue of CCC, suggesting their involvement in cardiomyopathy.
Discussion: Our findings highlight the ability of CD4+ T-cells from CCC patients to recognize and react to foreign and self-peptides, thereby emphasizing the importance of HLA-DRB1 alleles in the presentation of potentially pathogenic antigens and in the amplification of CCC pathology.
Keywords: Chagas disease; HLA-DRB1; T-cells; cardiomyopathy; cross-reactivity.
Copyright © 2025 Souza-Silva, Neves, Teixeira-Carvalho, Figueiredo, Morais, Apostólico, Rodrigues, Kalil, Juliano, Juliano, Araújo, Pantaleao, Mutarelli, Nunes, Gollob and Dutra.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures






Similar articles
-
Signatures of CD4+ T and B cells are associated with distinct stages of chronic chagasic cardiomyopathy.Front Immunol. 2024 Apr 25;15:1385850. doi: 10.3389/fimmu.2024.1385850. eCollection 2024. Front Immunol. 2024. PMID: 38726014 Free PMC article.
-
In Silico Guided Discovery of Novel Class I and II Trypanosoma cruzi Epitopes Recognized by T Cells from Chagas' Disease Patients.J Immunol. 2020 Mar 15;204(6):1571-1581. doi: 10.4049/jimmunol.1900873. Epub 2020 Feb 14. J Immunol. 2020. PMID: 32060134
-
Autoimmunity in Chagas' disease. Identification of cardiac myosin-B13 Trypanosoma cruzi protein crossreactive T cell clones in heart lesions of a chronic Chagas' cardiomyopathy patient.J Clin Invest. 1996 Oct 15;98(8):1709-12. doi: 10.1172/JCI118969. J Clin Invest. 1996. PMID: 8878420 Free PMC article.
-
Myocardial gene and protein expression profiles after autoimmune injury in Chagas' disease cardiomyopathy.Autoimmun Rev. 2011 Jan;10(3):163-5. doi: 10.1016/j.autrev.2010.09.019. Epub 2010 Sep 29. Autoimmun Rev. 2011. PMID: 20883825 Review.
-
Trypanosoma cruzi-induced molecular mimicry and Chagas' disease.Curr Top Microbiol Immunol. 2005;296:89-123. doi: 10.1007/3-540-30791-5_6. Curr Top Microbiol Immunol. 2005. PMID: 16323421 Review.
References
-
- Hamrouni S, Bras-Gonçalves R, Kidar A, Aoun K, Chamakh-Ayari R, Petitdidierid E, et al. . Design of multi-epitope peptides containing hla class-i and class-ii-restricted epitopes derived from immunogenic Leishmania proteins, and evaluation of CD4+ and CD8+ T cell responses induced in cured cutaneous leishmaniasis subjects. PloS Negl Trop Dis. (2020) 14:1–31. doi: 10.1371/journal.pntd.0008093 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

