Prediction of bloodstream infection using machine learning based primarily on biochemical data
- PMID: 40394073
- PMCID: PMC12092749
- DOI: 10.1038/s41598-025-01821-6
Prediction of bloodstream infection using machine learning based primarily on biochemical data
Abstract
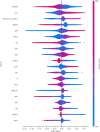
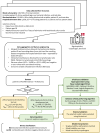
Early diagnosis of bloodstream infection (BSI) is crucial for informed antibiotic use. This study developed a machine learning approach for early BSI detection using a comprehensive dataset from Rigshospitalet, Denmark (2010-2020). The dataset included 144,398 samples from adult patients, containing blood culture results, demographics, and up to 36 biochemical variables. Positive blood culture was observed in 6.4% of samples, mostly caused by Staphylococcus aureus, Escherichia coli, and Enterococcus faecium. 80% of the samples (N = 43,351 patients) were used for ML model development and five-fold cross-validation, with 20% for independent testing (N = 10,837). Among seven models, LightGBM performed best, achieving an AUC of 0.69 on the test set. It was more accurate in detecting negatives, with a negative predictive value (NPV) of 0.96 and specificity of 0.74, compared to a positive predictive value (PPV) of 0.13 and sensitivity of 0.54. SHapley Additive exPlanations (SHAP) identified platelets, leukocytes, and neutrophils-to-lymphocytes as the top-3 predictive features. The model showed higher sensitivity (average 0.66) for common pathogens, e.g., 0.71 for E. coli. Results highlight the potential of biochemical variables as diagnostic factors for BSI, indicating clinical use to focus on identifying patients at low risks and can be further enhanced in future investigations.
Keywords: Artificial intelligence; Biomarkers; Classification models; Clinical utility; Diagnosis; Electronic medical records; Infection management; Interpretability; Real-world data.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Competing interests: The authors declare no competing interests. Ethical approval and considerations.: There were no ethical concerns for this study. This was a retrospective, non-intervention study. All investigations were performed retrospectively and on anonymized data in accordance with Danish guidelines and regulations. No human tissue was stored or used. All methods were carried out in accordance with relevant guidelines and regulations and the study approved by relevant agencies (see below). Due to the retrospective nature of the study, The Scientific Ethics Committee for the Capital Region of Denmark waived the need of obtaining informed consent. The extraction of data from registries was approved by the local data protection agency (Journal-nr.: R-21005123) and approved by the Danish Patient Safety Authority (journal no. 2021 -328).
Figures

Similar articles
-
Development and application of an early prediction model for risk of bloodstream infection based on real-world study.BMC Med Inform Decis Mak. 2025 May 14;25(1):186. doi: 10.1186/s12911-025-03020-9. BMC Med Inform Decis Mak. 2025. PMID: 40369550 Free PMC article.
-
Machine learning-based prognostic model for bloodstream infections in hematological malignancies using Th1/Th2 cytokines.BMC Infect Dis. 2025 Mar 26;25(1):415. doi: 10.1186/s12879-025-10808-7. BMC Infect Dis. 2025. PMID: 40140749 Free PMC article.
-
Enhancing pathogens detection in suspected geriatric bloodstream infections using Nanopore-targeted sequencing.Microbiol Spectr. 2025 Jan 7;13(1):e0155424. doi: 10.1128/spectrum.01554-24. Epub 2024 Nov 22. Microbiol Spectr. 2025. PMID: 39576187 Free PMC article.
-
Predicting community acquired bloodstream infection in infants using full blood count parameters and C-reactive protein; a machine learning study.Eur J Pediatr. 2024 Jul;183(7):2983-2993. doi: 10.1007/s00431-024-05441-6. Epub 2024 Apr 18. Eur J Pediatr. 2024. PMID: 38634890 Free PMC article.
-
Bloodstream infections in HIV-infected patients.Virulence. 2016 Apr 2;7(3):320-8. doi: 10.1080/21505594.2016.1158359. Virulence. 2016. PMID: 26950194 Free PMC article. Review.
References
-
- Kontula, K. S. K., Skogberg, K., Ollgren, J., Järvinen, A. & Lyytikäinen, O. Early deaths in bloodstream infections: a population-based case series. Infect. Dis. 48 (5), 379–385 (2016). - PubMed
-
- Papadimitriou-Olivgeris, M. et al. Predictors for delayed antibiotic administration among bacteraemic patients in the emergency department: differences between medical and surgical interns. Eur. J. Clin. Investig. 50 (11), 1–7 (2020). - PubMed
-
- Townsend, S. R. Antibiotic administration and timing: risks, delay, zombies**. Crit. Care Med. 49 (10), 1818–1821 (2021). - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

