Trajectories from single-cells to PAX5-driven leukemia reveal PAX5-MYC interplay in vivo
- PMID: 40394211
- PMCID: PMC12208885
- DOI: 10.1038/s41375-025-02626-2
Trajectories from single-cells to PAX5-driven leukemia reveal PAX5-MYC interplay in vivo
Abstract
PAX5 acts as a master regulator of B-cell proliferation and differentiation. Its germline and somatic deregulation have both been implicated in the development of B-cell precursor acute lymphoblastic leukemia (BCP-ALL). However, the process how reduced PAX5 transcriptional activity mediates progression to BCP-ALL, is still poorly understood. Here, we characterized the longitudinal effects of PAX5 reduction on healthy, pre-leukemic and BCP-ALL cells at the single-cell level. Cell-surface marker analysis revealed a genotype-driven enrichment of the pre-BII population in healthy Pax5± mice. This population showed downregulated B-cell receptor signaling, while DNA replication/repair and cell-cycle signaling pathways were upregulated. Moreover, we observed a shift in the kappa/lambda light chain ratio toward lambda rearranged B-cells. Transplantation experiments further validated a delay of Pax5± pre-BII cells in maturation and transition to IgM-positivity. Additionally, single-cell RNA-Sequencing and bulk ATAC-Sequencing of different stages of BCP-ALL evolution showed that Pax5± pre-leukemic cells lose their B-cell identity and display Myc activation. Subsequently, BCP-ALLs acquired additional RAG-mediated aberrations and driver mutations in JAK-STAT and RAS-signaling pathways. Together, this study elucidates molecular and functional checkpoints in PAX5-mediated pre-leukemic cell progression exploitable for therapeutic intervention and demonstrates that PAX5 reduction is sufficient to initiate clonal evolution to BCP-ALL through activation of MYC.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no potential conflicts of interest.
Figures



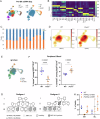



References
-
- Cobaleda C, Schebesta A, Delogu A, Busslinger M. Pax5: the guardian of B cell identity and function. Nat Immunol. 2007;8:463–70. - PubMed
-
- Nutt SL, Heavey B, Rolink AG, Busslinger M. Commitment to the B-lymphoid lineage depends on the transcription factor Pax5. Nature. 1999;401:556–62. - PubMed
-
- Fedl AS, Tagoh H, Gruenbacher S, Sun Q, Schenk RL, Froussios K, et al. Transcriptional function of E2A, Ebf1, Pax5, Ikaros and Aiolos analyzed by in vivo acute protein degradation in early B cell development. Nat Immunol. 2024;25:1663–77. - PubMed
MeSH terms
Substances
Grants and funding
- Stg 85222/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 European Research Council (H2020 Excellent Science - European Research Council)
- 70114539/Deutsche Krebshilfe (German Cancer Aid)
- 547295412/Deutsche Forschungsgemeinschaft (German Research Foundation)
- DJCLS 18R/2021/José Carreras Leukämie-Stiftung (Deutsche José Carreras Leukämie-Stiftung)
- 341693/Sigrid Juséliuksen Säätiö (Sigrid Jusélius Foundation)
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

