AP5Z1 affects hepatocellular carcinoma growth and autophagy by regulating PTEN ubiquitination and modulating the PI3K/Akt/mTOR pathway
- PMID: 40394639
- PMCID: PMC12090622
- DOI: 10.1186/s12967-025-06537-9
AP5Z1 affects hepatocellular carcinoma growth and autophagy by regulating PTEN ubiquitination and modulating the PI3K/Akt/mTOR pathway
Abstract
Background: Hepatocellular carcinoma (HCC) is a leading cause of cancer death worldwide, with high incidence and mortality rates, and the number of cases is expected to increase by 2030. Understanding the molecular mechanisms of HCC and identifying new therapeutic targets and biomarkers for HCC are crucial.
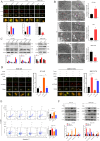
Methods: In this study, we examined adaptor-related protein complex 5 subunit ζ1 (AP5Z1) expression in liver cancer and nearby noncancerous tissues to explore its effects on HCC cell growth, death, and autophagy. The functional and molecular mechanisms of AP5Z1 were studied using clinical sample analysis, Western blot (WB), immunohistochemistry (IHC), quantitative reverse-transcription polymerase chain reaction (qRT‒PCR), coimmunoprecipitation (Co-IP), cell proliferation assays, flow cytometry (FCM), autophagy assays, electron microscopy, mass spectrometry (MS), transcriptome analysis, and animal model experiments.
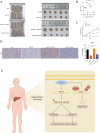
Results: AP5Z1 expression was notably higher in HCC tissues than in normal tissues and was linked to a poor prognosis. The results of both in vitro and in vivo studies revealed that AP5Z1 promoted HCC cell growth and reduced apoptosis. In addition, AP5Z1 regulates cellular autophagy by ubiquitinating the phosphatase and tensin homolog (PTEN) protein and modulating the phosphoinositide 3-kinase (PI3K)/protein kinase B (Akt)/mammalian target of rapamycin (mTOR) pathway.
Conclusions: AP5Z1 influences autophagy and apoptosis in HCC cells by interacting with PTEN to modulate the PI3K/Akt/mTOR pathway. This gene might promote PTEN ubiquitination and degradation by recruiting tripartite motif-containing protein 21 (TRIM21), making it a potential biomarker for diagnosing and predicting the outcome of HCC as well as a target for new treatment strategies.
Keywords: AP5Z1; Apoptosis; Autophagy; Hepatocellular carcinoma; PI3K/Akt/mTOR signalling pathway; PTEN; TRIM21; Ubiquitination.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: The study was approved by the Committee on the Ethics of the First Affiliated Hospital of Guangxi Medical University (Approval number: 2024-E754-01) on October 24, 2024. Additionally, the animal experiments were approved by the Laboratory Animal Ethics Committee of Guangxi Medical University, with the ethics number: No. 202405005. Consent for publication: Written informed consent for publication was obtained from all participants. Completing interest: All authors declared no competing interest.
Figures




Similar articles
-
Alpha1-ACT Functions as a Tumour Suppressor in Hepatocellular Carcinoma by Inhibiting the PI3K/AKT/mTOR Signalling Pathway via Activation of PTEN.Cell Physiol Biochem. 2017;41(6):2289-2306. doi: 10.1159/000475648. Epub 2017 Apr 26. Cell Physiol Biochem. 2017. PMID: 28456796
-
MicroRNA-155-5p promotes hepatocellular carcinoma progression by suppressing PTEN through the PI3K/Akt pathway.Cancer Sci. 2017 Apr;108(4):620-631. doi: 10.1111/cas.13177. Epub 2017 Apr 19. Cancer Sci. 2017. PMID: 28132399 Free PMC article.
-
Down-regulated lncRNA TP73-AS1 reduces radioresistance in hepatocellular carcinoma via the PTEN/Akt signaling pathway.Cell Cycle. 2019 Nov;18(22):3177-3188. doi: 10.1080/15384101.2019.1671089. Epub 2019 Sep 29. Cell Cycle. 2019. Retraction in: Cell Cycle. 2023 Jul-Aug;22(14-16):1802. doi: 10.1080/15384101.2023.2211863. PMID: 31564201 Free PMC article. Retracted.
-
Role of regulatory miRNAs of the PI3K/AKT/mTOR signaling in the pathogenesis of hepatocellular carcinoma.J Cell Physiol. 2020 May;235(5):4146-4152. doi: 10.1002/jcp.29333. Epub 2019 Oct 29. J Cell Physiol. 2020. PMID: 31663122 Review.
-
Research on the role and mechanism of the PI3K/Akt/mTOR signalling pathway in osteoporosis.Front Endocrinol (Lausanne). 2025 May 12;16:1541714. doi: 10.3389/fendo.2025.1541714. eCollection 2025. Front Endocrinol (Lausanne). 2025. PMID: 40421249 Free PMC article. Review.
References
-
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019;69:7–34. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

