Drug repurposing candidates for amyotrophic lateral sclerosis using common and rare genetic variants
- PMID: 40395632
- PMCID: PMC12089939
- DOI: 10.1093/braincomms/fcaf184
Drug repurposing candidates for amyotrophic lateral sclerosis using common and rare genetic variants
Abstract
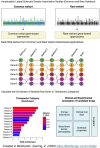
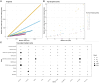
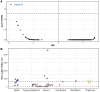
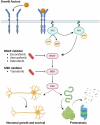
Amyotrophic lateral sclerosis (ALS) is a devastating neurodegenerative condition for which novel disease modifying therapies are urgently needed. Given the increasing bottlenecks in drug discovery pipelines, repurposing existing drugs for ALS may represent a path to expedite translation and improve disease outcomes. However, ALS is a heterogeneous disease for which the aetiology remains poorly characterized, complicating efforts to effectively repurpose drugs. We propose that the polygenic architecture of ALS genetic liability, which ranges from ultra-rare, high-impact variation to common frequency loci of small-individual effect, could be leveraged to prioritize drug repurposing candidates which are more generalizable to the ALS clinical population. Here, we utilize common and rare frequency ALS genetic risk with a novel approach to uncover therapeutic classes that may be prospective repurposing opportunities in ALS. The common variant-led analyses integrated both positional-based and functional gene-based tests on SNP-genotype data from a genome-wide association study of ALS and implicated mitogen-activated protein kinase signalling related downregulation through B-Raf inhibitors as a prospective target for repurposing. The rare variant-led approaches leveraged rare variant burden testing of exonic variation on whole genome-sequencing data from a subset of the common variant genome-wide association study cohort and prioritized B-vitamin related candidates, such as cobalamin and niacin. Clinical characterization of these putative repurposing opportunities revealed genetic support to existing biology for which related compounds are actively proceeding through ALS clinical studies. Moreover, leveraging transcriptomic data from ALS derived cell lines carrying a selection of pathogenic variants in genes that cause familial forms of ALS (C9orf72, SOD1, FUS and TARDBP) suggested that the action of B-Raf inhibitors may be of particular relevance to C9orf72 carriers, whilst the signal for B-vitamin signalling related targets was strongest in SOD1 carriers. In summary, we demonstrate the importance of considering the therapeutic actionability of both common and rare-variant mediated risk for ALS given the immense biological heterogeneity of this disorder. Future pre-clinical and clinical studies are now warranted to further characterize the tractability of these prioritized compounds.
Keywords: drug repurposing; exome-wide association studies; genome-wide association studies; motor neurone disease; stem cells.
© The Author(s) 2025. Published by Oxford University Press on behalf of the Guarantors of Brain.
Conflict of interest statement
W.R.R. discloses equity held in a genetics company (PolygenRx Pty. Ltd.) unrelated to this work. The remaining authors declare no competing financial interests.
Figures


References
-
- Hardiman O, Al-Chalabi A, Chio A, et al. Amyotrophic lateral sclerosis. Nat Rev Dis Primer. 2017;3:17071. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
