Time-Lapse Super-Resolution Imaging and Optical Manipulation of Growth Cones in Elongating Axons and Migrating Neurons
- PMID: 40395842
- PMCID: PMC12086313
- DOI: 10.21769/BioProtoc.5251
Time-Lapse Super-Resolution Imaging and Optical Manipulation of Growth Cones in Elongating Axons and Migrating Neurons
Abstract
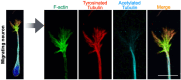
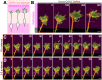
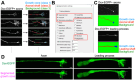
The growth cone is a highly motile tip structure that guides axonal elongation and directionality in differentiating neurons. Migrating immature neurons also exhibit a growth cone-like structure (GCLS) at the tip of the leading process. However, it remains unknown whether the GCLS in migrating immature neurons shares the morphological and molecular features of axonal growth cones and can thus be considered equivalent to them. Here, we describe a detailed method for time-lapse imaging and optical manipulation of growth cones using a super-resolution laser-scanning microscope. To observe growth cones in elongating axons and migrating neurons, embryonic cortical neurons and neonatal ventricular-subventricular zone (V-SVZ)-derived neurons, respectively, were transfected with plasmids encoding fluorescent protein-conjugated cytoskeletal probes and three-dimensionally cultured in Matrigel, which mimics the in vivo background. At 2-5 days in vitro, the morphology and dynamics of these growth cones and their associated cytoskeletal molecules were assessed by time-lapse super-resolution imaging. The use of photoswitchable cytoskeletal inhibitors, which can be reversibly and precisely controlled by laser illumination at two different wavelengths, revealed the spatiotemporal regulatory machinery and functional significance of growth cones in neuronal migration. Furthermore, machine learning-based methods enabled us to automatically segment growth cone morphology from elongating axons and the leading process. This protocol provides a cutting-edge methodology for studying the growth cone in developmental and regenerative neuroscience, being adaptable for various cell biology and imaging applications. Key features • Three-dimensional primary culture of migrating and differentiating neurons in Matrigel. • Visualization of fine morphology and dynamics of growth cones using super-resolution imaging. • Optical manipulation of cytoskeletal molecules in growth cones using photoswitchable inhibitors. • Machine learning-based extraction of growth cone morphology.
Keywords: Elongating axons; F-actin; Growth cone; Microtubules; Migrating neurons; Optical manipulation; Photoswitchable inhibitor; Postnatal neurogenesis; Super-resolution imaging; Ventricular–subventricular zone.
©Copyright : © 2025 The Authors; This is an open access article under the CC BY-NC license.
Conflict of interest statement
Competing interestsThe authors declare no competing interests.
Figures


Similar articles
-
The Difference in the Cytoskeletal Machinery of Growth Cones of Growing Axons and Leading Processes.Dev Neurosci. 2022;44(6):455-465. doi: 10.1159/000522200. Epub 2022 Jan 31. Dev Neurosci. 2022. PMID: 35100584
-
The Microtubule-Associated Protein Tau Mediates the Organization of Microtubules and Their Dynamic Exploration of Actin-Rich Lamellipodia and Filopodia of Cortical Growth Cones.J Neurosci. 2018 Jan 10;38(2):291-307. doi: 10.1523/JNEUROSCI.2281-17.2017. Epub 2017 Nov 22. J Neurosci. 2018. PMID: 29167405 Free PMC article.
-
Identification of the growth cone as a probe and driver of neuronal migration in the injured brain.Nat Commun. 2024 Mar 9;15(1):1877. doi: 10.1038/s41467-024-45825-8. Nat Commun. 2024. PMID: 38461182 Free PMC article.
-
Drebrin in Neuronal Migration and Axonal Growth.Adv Exp Med Biol. 2017;1006:141-155. doi: 10.1007/978-4-431-56550-5_9. Adv Exp Med Biol. 2017. PMID: 28865019 Review.
-
Directional guidance of nerve growth cones.Curr Opin Neurobiol. 2006 Feb;16(1):52-8. doi: 10.1016/j.conb.2005.12.005. Epub 2006 Jan 4. Curr Opin Neurobiol. 2006. PMID: 16387488 Review.
References
-
- Stoeckli E. T.(2018). Understanding axon guidance: are we nearly there yet? Development 145(10): e151415. https://doi.org/10.1242/dev.151415 - DOI - PubMed
-
- Cajal, S. R.(1890). A quelle époque apparaissent les expansions des cellules nerveuses de la moelle épinière du poulet. Anat Anz. 5: 609–613.
LinkOut - more resources
Full Text Sources

