Enhancing photosynthesis under salt stress via directed evolution in cyanobacteria
- PMID: 40397698
- PMCID: PMC12127731
- DOI: 10.1093/plphys/kiaf209
Enhancing photosynthesis under salt stress via directed evolution in cyanobacteria
Abstract
A key aspect of enhancing photosynthesis is improving the kinetics of photochemical quenching recovery following environmental perturbation or stress. Salt stress exacerbates high light stress in cyanobacteria and leads to severe yield losses in crop plants. Genetic traits that confer salt tolerance without compromising photosynthetic performance are essential for improving photosynthesis under these conditions. Here, we applied accelerated evolution in Synechococcus elongatus PCC 7942 by conditionally suppressing its methyl-directed mismatch repair system to obtain beneficial genetic traits for enhanced photosynthesis under salt stress. We screened over 10,000 mutants and isolated 8 strains with increased biomass or sucrose productivity under salt stress. Genome sequencing revealed an average of 8 to 20 single nucleotide polymorphisms or indels per genome. Notably, mutations in the photosystem II (PSII) reaction center D1-encoding gene, resulting in the amino acid changes L353F, I358N, and H359N at the carboxyl terminus of the precursor-D1 (pD1) protein, improved photosynthesis under salt and combined salt and light stress by potentially accelerating D1 maturation during PSII repair. Phylogenetic analysis of pD1 across cyanobacteria and red algae highlights the broad relevance of these adaptive genetic traits, underscoring the importance of leveraging evolutionary insights to improve photosynthesis under stress or fluctuating environments.
© The Author(s) 2025. Published by Oxford University Press on behalf of American Society of Plant Biologists.
Conflict of interest statement
Conflict of interest statement. X.W. and Z.J. are listed as inventors in a patent application (application no. 18/400,586) for applying the high-throughput screening method to uncover genetic traits for photosynthesis improvement. X.W. is listed as the inventor for another provisional patent application (application no. 63/655,889) to apply pD1 mutations to improve photosynthesis and plant growth. The remaining authors declare no competing interests.
Figures




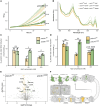

References
-
- Allakhverdiev SI, Nishiyama Y, Miyairi S, Yamamoto H, Inagaki N, Kanesaki Y, Murata N. Salt stress inhibits the repair of photodamaged photosystem II by suppressing the transcription and translation of psbA genes in Synechocystis. Plant Physiol. 2002:130(3):1443–1453. 10.1104/pp.011114 - DOI - PMC - PubMed
-
- Andrews S. FastQC: a quality control tool for high throughput sequence data. Cambridge, UK: Babraham Bioinformatics; 2010. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources

