Metaproteomics in the One Health framework for unraveling microbial effectors in microbiomes
- PMID: 40410872
- PMCID: PMC12100821
- DOI: 10.1186/s40168-025-02119-5
Metaproteomics in the One Health framework for unraveling microbial effectors in microbiomes
Abstract
One Health seeks to integrate and balance the health of humans, animals, and environmental systems, which are intricately linked through microbiomes. These microbial communities exchange microbes and genes, influencing not only human and animal health but also key environmental, agricultural, and biotechnological processes. Preventing the emergence of pathogens as well as monitoring and controlling the composition of microbiomes through microbial effectors including virulence factors, toxins, antibiotics, non-ribosomal peptides, and viruses holds transformative potential. However, the mechanisms by which these microbial effectors shape microbiomes and their broader functional consequences for host and ecosystem health remain poorly understood. Metaproteomics offers a novel methodological framework as it provides insights into microbial dynamics by quantifying microbial biomass composition, metabolic functions, and detecting effectors like viruses, antimicrobial resistance proteins, and non-ribosomal peptides. Here, we highlight the potential of metaproteomics in elucidating microbial effectors and their impact on microbiomes and discuss their potential for modulating microbiomes to foster desired functions.
Keywords: Metaproteomics; Microbial community; Bacteriophages; Microbiome; Microbial effectors.
© 2025. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures


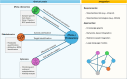

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

