Selected Alu methylation levels in the gastric carcinogenesis cascade
- PMID: 40416611
- PMCID: PMC12101442
- DOI: 10.7717/peerj.19485
Selected Alu methylation levels in the gastric carcinogenesis cascade
Abstract
Background: Genome-wide hypomethylation, a common epigenetic change that occurs during cancer development, primarily affects repetitive elements, such as Alu repeats. Consequently, Alu repeats can be used as a surrogate marker of genomic hypomethylation.
Methods: In this study, we aimed to investigate the correlation between Alu methylation levels and the multistage course of gastric carcinogenesis.
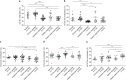
Results: We found that the Alu methylation levels in gastric cancer tissue decreased compared with those in normal gastric tissue, with the change in methylation levels and pattern being most significant between chronic gastritis and intestinal metaplasia. Moreover, Alu methylation levels were not associated with Helicobacter pylori or Epstein-Barr virus infection.
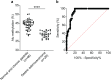
Conclusions: Finally, our sensitivity and specificity analyses suggested that Alu methylation level can be used to distinguish gastric cancer tissue from normal tissue. Thus, Alu methylation level shows promise as biomarker for gastric cancer diagnosis.
Keywords: Epstein–Barr virus; Gastric cancer; Helicobacter pylori; Hypomethylation.
© 2025 Meevassana et al.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures

Similar articles
-
Alu and Satα hypomethylation in Helicobacter pylori-infected gastric mucosae.Int J Cancer. 2011 Jan 1;128(1):33-9. doi: 10.1002/ijc.25534. Int J Cancer. 2011. PMID: 20602342
-
ALU and LINE-1 hypomethylations in multistep gastric carcinogenesis and their prognostic implications.Int J Cancer. 2012 Sep 15;131(6):1323-31. doi: 10.1002/ijc.27369. Epub 2012 Jan 3. Int J Cancer. 2012. PMID: 22120154
-
Comparison of CpG island hypermethylation and repetitive DNA hypomethylation in premalignant stages of gastric cancer, stratified for Helicobacter pylori infection.J Pathol. 2009 Dec;219(4):410-6. doi: 10.1002/path.2596. J Pathol. 2009. PMID: 19639607
-
Helicobacter pylori-induced inflammation and epigenetic changes during gastric carcinogenesis.World J Gastroenterol. 2015 Dec 7;21(45):12742-56. doi: 10.3748/wjg.v21.i45.12742. World J Gastroenterol. 2015. PMID: 26668499 Free PMC article. Review.
-
DNA methylation in gastric cancer, related to Helicobacter pylori and Epstein-Barr virus.World J Gastroenterol. 2014 Apr 14;20(14):3916-26. doi: 10.3748/wjg.v20.i14.3916. World J Gastroenterol. 2014. PMID: 24744581 Free PMC article. Review.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

