Duplication of the autism-related gene Chd8 leads to behavioral hyperactivity and neurodevelopmental defects in mice
- PMID: 40419468
- PMCID: PMC12106843
- DOI: 10.1038/s41467-025-59853-5
Duplication of the autism-related gene Chd8 leads to behavioral hyperactivity and neurodevelopmental defects in mice
Abstract
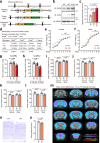
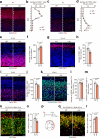
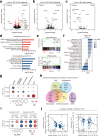
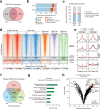
Mutations in the gene encoding chromodomain helicase DNA-binding protein 8 (CHD8) are strongly associated with autism spectrum disorder (ASD). Although duplications of the chromosomal locus including CHD8 have also been detected in individuals with neurodevelopmental disorders, the contribution of CHD8 duplication to clinical phenotypes and the underlying mechanisms have remained unknown. Here we show that Chd8 knock-in (KI) mice that overexpress CHD8 as a model of human CHD8 duplication manifest growth retardation, microcephaly, impaired neuronal differentiation, and behavioral abnormalities including hyperactivity and reduced anxiety-like behavior. Chd8 overexpression affects the transcription and chromatin accessibility of genes related to neurogenesis, with these changes being associated with aberrant binding of CHD8 to enhancer regions. Furthermore, pharmacological intervention partially ameliorates the hyperactivity of Chd8 KI mice. Our results thus indicate that Chd8 KI mice recapitulate key features of CHD8 duplication syndrome in humans, providing insight into pathogenic mechanisms underlying neurodevelopmental disorders.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures
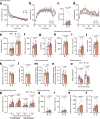
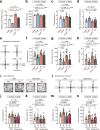
Similar articles
-
Oligodendrocyte dysfunction due to Chd8 mutation gives rise to behavioral deficits in mice.Hum Mol Genet. 2020 May 28;29(8):1274-1291. doi: 10.1093/hmg/ddaa036. Hum Mol Genet. 2020. PMID: 32142125
-
The Deficiency of the ASD-Related Gene CHD8 Disrupts Behavioral Patterns and Inhibits Hippocampal Neurogenesis in Mice.J Mol Neurosci. 2024 Oct 31;74(4):103. doi: 10.1007/s12031-024-02283-7. J Mol Neurosci. 2024. PMID: 39480606
-
Heterozygous deletion of the autism-associated gene CHD8 impairs synaptic function through widespread changes in gene expression and chromatin compaction.Am J Hum Genet. 2023 Oct 5;110(10):1750-1768. doi: 10.1016/j.ajhg.2023.09.004. Am J Hum Genet. 2023. PMID: 37802044 Free PMC article.
-
The Mechanisms of CHD8 in Neurodevelopment and Autism Spectrum Disorders.Genes (Basel). 2021 Jul 26;12(8):1133. doi: 10.3390/genes12081133. Genes (Basel). 2021. PMID: 34440307 Free PMC article. Review.
-
Neurodevelopmental functions of CHD8: new insights and questions.Biochem Soc Trans. 2024 Feb 28;52(1):15-27. doi: 10.1042/BST20220926. Biochem Soc Trans. 2024. PMID: 38288845 Free PMC article. Review.
References
-
- Parenti, I., Rabaneda, L. G., Schoen, H. & Novarino, G. Neurodevelopmental disorders: from genetics to functional pathways. Trends Neurosci.43, 608–621 (2020). - PubMed
-
- Cook, E. H. Jr & Scherer, S. W. Copy-number variations associated with neuropsychiatric conditions. Nature455, 919–923 (2008). - PubMed
-
- Grayton, H. M., Fernandes, C., Rujescu, D. & Collier, D. A. Copy number variations in neurodevelopmental disorders. Prog. Neurobiol.99, 81–91 (2012). - PubMed
MeSH terms
Substances
Grants and funding
- JP21H02847/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP24H00627/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP21J00911/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP21K15726/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP21H05619/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP22H05493/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP23K14795/MEXT | Japan Society for the Promotion of Science (JSPS)
- JP22gm6310008/Japan Agency for Medical Research and Development (AMED)
- JP24wm0625316/Japan Agency for Medical Research and Development (AMED)
- JP24wm0625001/Japan Agency for Medical Research and Development (AMED)
- JP24wm0625308/Japan Agency for Medical Research and Development (AMED)
- JP24wm0625308/Japan Agency for Medical Research and Development (AMED)
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

