Transcriptomic and Proteomic Profiling of Rabbit Kidney Cells Infected with Equine Herpesvirus 8
- PMID: 40431657
- PMCID: PMC12115596
- DOI: 10.3390/v17050647
Transcriptomic and Proteomic Profiling of Rabbit Kidney Cells Infected with Equine Herpesvirus 8
Abstract
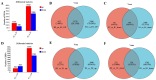
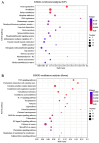
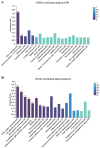
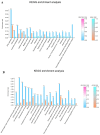
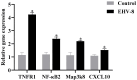
The present study investigated the host cell response to EHV-8 infection in rabbit kidney (RK-13) cells through transcriptomic and proteomic approaches. At 24 h post-infection, a total of 2118 differentially expressed genes (DEGs) were identified, with 1338 upregulated and 780 downregulated. At 48 h, 7388 DEGs were detected, with 4342 upregulated and 3046 downregulated genes. Proteomic analysis revealed 932 differentially expressed proteins (DEPs) at 24 h (364 upregulated and 568 downregulated) and 3866 DEPs at 48 h (2285 upregulated and 1581 downregulated). Of these, 237 upregulated and 336 downregulated proteins were common across both time points. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis indicated that the majority of DEGs and DEPs were enriched in key inflammation-related pathways, notably the TNF and NF-κB signaling pathways. Validation of the transcriptomic and proteomic data was performed using RT-PCR and parallel reaction monitoring (PRM), respectively, and confirmed consistent trends for TNFR1, NF-κB p65, and MAP3K8, as reported in the transcriptomic and proteomic screening. These findings suggest that EHV-8 infection may modulate host immune responses by activating the TNF signaling pathway. However, given that RK-13 cells may not fully replicate viral-host interactions in equine species, further in vivo studies in horses and donkeys are required to provide a more comprehensive understanding of the viral pathogenesis in these animals.
Keywords: EHV-8; RK-13 cells; TNF signaling pathway; immune response; proteomics; transcriptomics.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

Similar articles
-
CXCL14 drives age-related intervertebral disc degeneration via NF-κB pathway activation in a multiomic study.Sci Rep. 2025 Jul 13;15(1):25307. doi: 10.1038/s41598-025-10998-9. Sci Rep. 2025. PMID: 40653514 Free PMC article.
-
Identification of shared key genes and pathways in osteoarthritis and sarcopenia patients based on bioinformatics analysis.Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2025 Mar 28;50(3):430-446. doi: 10.11817/j.issn.1672-7347.2025.240669. Zhong Nan Da Xue Xue Bao Yi Xue Ban. 2025. PMID: 40628511 Chinese, English.
-
Single-cell proteomics analysis of human oocytes during GV-to-MI transition.Hum Reprod. 2025 Jul 1;40(7):1332-1343. doi: 10.1093/humrep/deaf086. Hum Reprod. 2025. PMID: 40359387
-
The effect of sample site and collection procedure on identification of SARS-CoV-2 infection.Cochrane Database Syst Rev. 2024 Dec 16;12(12):CD014780. doi: 10.1002/14651858.CD014780. Cochrane Database Syst Rev. 2024. PMID: 39679851 Free PMC article.
-
Intravenous magnesium sulphate and sotalol for prevention of atrial fibrillation after coronary artery bypass surgery: a systematic review and economic evaluation.Health Technol Assess. 2008 Jun;12(28):iii-iv, ix-95. doi: 10.3310/hta12280. Health Technol Assess. 2008. PMID: 18547499
References
-
- Pusterla N., Leutenegger C.M., Wilson W.D., Watson J.L., Ferraro G.L., Madigan J.E. Equine Herpesvirus-4 Kinetics in Peripheral Blood Leukocytes and Nasopharyngeal Secretions in Foals Using Quantitative Real-Time TaqMan PCR. J. Vet. Diagn. Investig. 2005;17:578–581. doi: 10.1177/104063870501700610. - DOI - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

