Skin microbiomes of frogs vary among body regions, revealing differences that reflect known patterns of chytrid infection
- PMID: 40432964
- PMCID: PMC12106533
- DOI: 10.3389/fmicb.2025.1579231
Skin microbiomes of frogs vary among body regions, revealing differences that reflect known patterns of chytrid infection
Abstract
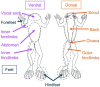
Introduction: The amphibian skin microbiome is an important line of defense against pathogens including the deadly chytrid fungus, Batrachochytrium dendrobatidis (Bd). Bd is known to preferentially infect ventral skin surfaces and feet of host amphibians, often leaving dorsal surfaces like the back uninfected. Within-individual variation in infection distribution across the skin, therefore, may relate to differences in microbiomes among skin regions. However, microbiome heterogeneity within amphibian individuals remains poorly characterized.
Methods: We utilized 16S rRNA gene amplicon sequencing to compare microbiomes of 10 body regions from nine captive Rana sierrae individuals and their tank environments. These individuals were naive to Bd, allowing us to assess whether microbiomes differed among body regions prior to any impacts that may be caused by infection.
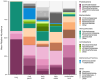
Results: We found that frog skin and tank environments harbored distinct microbial communities. On frog skin, the bacterial families Burkholderiaceae (phylum Proteobacteria) and Rubritaleaceae (phylum Verrucomicrobia) were dominant, driven in large part by relative abundances of undescribed members of these families that were significantly higher on frogs than in their environment. Within individuals, we detected differences between microbiomes of body regions where Bd infection would be expected compared to regions that infrequently experience infection. Notably, putative Bd-inhibitory relative abundance was significantly higher on body regions where Bd infection is often localized.
Discussion: These findings suggest that microbiomes in certain skin regions may be predisposed for interactions with Bd. Further, our results highlight the importance of considering intraindividual heterogeneities, which could provide insights relevant to predicting localized interactions with pathogens.
Keywords: Batrachochytrium dendrobatidis; Rana sierrae; amphibian; captivity; microbiome; skin; within-individual heterogeneity.
Copyright © 2025 Ghose and Eisen.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



Update of
-
Skin microbiomes of frogs vary among individuals and body regions, revealing differences that reflect known patterns of chytrid infection.bioRxiv [Preprint]. 2025 Feb 8:2025.02.05.636728. doi: 10.1101/2025.02.05.636728. bioRxiv. 2025. Update in: Front Microbiol. 2025 May 13;16:1579231. doi: 10.3389/fmicb.2025.1579231. PMID: 39975414 Free PMC article. Updated. Preprint.
References
-
- Anderson M. J. (2001). A new method for non parametric multivariate analysis of variance. Austral Ecol. 26, 32–46. doi: 10.1111/j.1442-9993.2001.01070.pp.x, PMID: - DOI
-
- Asangba A. E., Mugisha L., Rukundo J., Lewis R. J., Halajian A., Cortés-Ortiz L., et al. (2022). Large comparative analyses of primate body site microbiomes indicate that the oral microbiome is unique among all body sites and conserved among nonhuman primates. Microbiol. Spectr. 10:e0164321. doi: 10.1128/spectrum.01643-21, PMID: - DOI - PMC - PubMed

