Blood Biomarker-Based Predictive Indicator for Liver Metastasis in Alpha-Fetoprotein-Producing Gastric Cancer and Multi-Omics Tumor Microenvironment Insights
- PMID: 40433893
- PMCID: PMC12302549
- DOI: 10.1002/advs.202503499
Blood Biomarker-Based Predictive Indicator for Liver Metastasis in Alpha-Fetoprotein-Producing Gastric Cancer and Multi-Omics Tumor Microenvironment Insights
Abstract
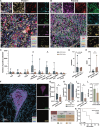
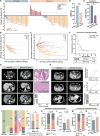
Alpha-fetoprotein-producing gastric cancer (AFPGC) is a rare but highly aggressive subtype of gastric cancer. Patients with AFPGC are at high risk of liver metastasis, and the tumor microenvironment (TME) is complex. A multicenter retrospective study is conducted from January 2011 to December 2021 and included 317 AFPGC patients. Using a multivariable logistic regression model, a nomogram for predicting liver metastasis is built. By combining AFP and the neutrophil-lymphocyte ratio (NLR), we developed a novel and easily applicable predictive indicator, termed ANLiM score, for liver metastasis in AFPGC. An integrated multi-omics analysis, including whole-exome sequencing and proteomic analysis, is conducted and revealed an immunosuppressive TME in AFPGC with liver metastasis. Single-cell RNA sequencing and multiplex immunofluorescence identified the potential roles of tumor-associated neutrophils and tertiary lymphoid structures in shaping the immune microenvironment. These findings are validated in a real-world cohort receiving anti-programmed cell death 1 (anti-PD-1) therapy, which showed concordant effectiveness. In addition, the ANLiM score is also identified as a promising biomarker for predicting immunotherapy efficacy. Overall, a blood biomarker-based predictive indicator is developed for liver metastasis and immunotherapy response in AFPGC. The findings on immune microenvironmental alterations for AFPGC with liver metastasis provide new insights for optimizing immunotherapy strategies.
Keywords: alpha‐fetoprotein‐producing gastric cancer; immunotherapy; liver metastasis; tumor microenvironment.
© 2025 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
