hnRNPM regulates influenza A virus replication through distinct mechanisms in human and avian cells: implications for cross-species transmission
- PMID: 40434105
- PMCID: PMC12172489
- DOI: 10.1128/jvi.00067-25
hnRNPM regulates influenza A virus replication through distinct mechanisms in human and avian cells: implications for cross-species transmission
Abstract
The eight-segmented RNA genome of influenza A virus (IAV) is transcribed and spliced into 10 major viral mRNAs in the nucleus of infected cells. Both transcription and splicing are facilitated by the host RNA polymerase II (Pol II) machinery via interactions between the viral ribonucleoprotein (vRNP) complex and various host factors. In this study, we demonstrate that IAV vRNPs recruit species-specific heterogeneous nuclear ribonucleoprotein M (hnRNPM) to support their replication in human and avian cells through distinct mechanisms. In A549 cells, human hnRNPM specifically facilitates the efficient transcription of HA, NA, M, and NS segments of WSN virus in a gene coding sequence-dependent manner. In contrast, in DF-1 cells, chicken hnRNPM restricts excessive splicing of M segment mRNA to ensure proper M2 protein production. Notably, human hnRNPM, with 34 additional amino acids compared with its chicken counterpart, fails to inhibit the M2 expression in DF-1 cells, whereas both human and chicken hnRNPM regulate WSN virus replication similarly in A549 cells. These findings highlight the host-specific roles of M2 levels in IAV replication and reveal how IAV co-opts host factors through virus genome sequence-dependent and host species-specific mechanisms, underscoring its high flexibility and adaptability during cross-species transmission.IMPORTANCEThe transcription and splicing of IAV genome in the nucleus of infected cells are precisely regulated to produce optimal amounts of viral proteins, ensuring efficient virus replication. In this study, we discovered that human hnRNPM regulates the IAV segment-specific differential transcription in a coding sequence-dependent manner in human cells. In contrast, chicken hnRNPM specifically inhibits M2 mRNA splicing to maintain proper M2 protein levels in avian cells. These species-specific regulatory mechanisms highlight the distinct replication strategies employed by IAV in human versus avian cells and underscore the complexity of cross-species transmission.
Keywords: M segment splicing; chicken hnRNPM; human hnRNPM; influenza A virus; segment-specific transcription.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


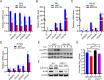


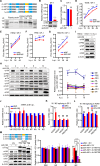


Similar articles
-
Viral polymerase-host interaction analysis reveals that the association between avian NUP93 and PB1 promotes H5N6 avian influenza virus replication.Microbiol Spectr. 2025 Jul;13(7):e0312024. doi: 10.1128/spectrum.03120-24. Epub 2025 May 23. Microbiol Spectr. 2025. PMID: 40407330 Free PMC article.
-
Comparative IP-MS Reveals HSPA5 and HSPA8 Interacting with Hemagglutinin Protein to Promote the Replication of Influenza A Virus.Pathogens. 2025 May 27;14(6):535. doi: 10.3390/pathogens14060535. Pathogens. 2025. PMID: 40559543 Free PMC article.
-
CRISPR editing of candidate host factors that impact influenza A virus infection.Microbiol Spectr. 2025 Mar 4;13(3):e0262724. doi: 10.1128/spectrum.02627-24. Epub 2025 Jan 31. Microbiol Spectr. 2025. PMID: 39887213 Free PMC article.
-
Host-Specific and Segment-Specific Evolutionary Dynamics of Avian and Human Influenza A Viruses: A Systematic Review.PLoS One. 2016 Jan 13;11(1):e0147021. doi: 10.1371/journal.pone.0147021. eCollection 2016. PLoS One. 2016. PMID: 26760775 Free PMC article.
-
Understanding the pandemic potential of the avian influenza Virus- key insights into pathogenesis, transmission, and host immunity.Arch Microbiol. 2025 Jun 23;207(8):185. doi: 10.1007/s00203-025-04371-7. Arch Microbiol. 2025. PMID: 40549164 Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

