Mechanistic divergence between SOS response activation and antibiotic-induced plasmid conjugation in Escherichia coli
- PMID: 40434128
- PMCID: PMC12211044
- DOI: 10.1128/spectrum.00090-25
Mechanistic divergence between SOS response activation and antibiotic-induced plasmid conjugation in Escherichia coli
Abstract
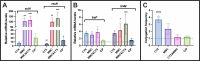
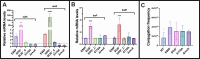
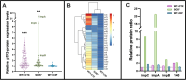
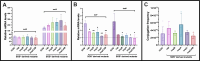
The SOS response is a critical DNA damage repair mechanism in bacteria, designed to counteract genotoxic stress and ensure survival. This system can be activated by different classes of antimicrobial agents, each inducing the SOS response through different mechanisms. Moreover, it has been observed that certain antibiotics can enhance conjugative plasmid transfer frequencies. However, while previous studies have suggested that the SOS response contributes to horizontal transfer of certain genes, its role in plasmid conjugation remains unclear. In this study, we investigated the relationship between the SOS response and conjugation of IncI1 and IncFII plasmids harboring various blaCTX-M resistance genes. Results showed that cefotaxime and mitomycin C induced both the SOS response and conjugation, while ciprofloxacin induced the SOS response without affecting conjugation frequencies. Further analysis of SOS mutants, ranging from constitutively inactive to hyper-induced states, revealed no correlation between SOS levels and conjugation frequencies, despite upregulation of tra gene expression in a SOS hyper-induced strain. Proteomic analysis revealed that cefotaxime-induced conjugation was associated with increased transfer and pilus protein expression. In contrast, the SOS hyper-induced strain displayed limited upregulation of plasmid-encoded proteins, suggesting post-transcriptional regulation. Additionally, putative LexA binding sites on the IncI1 plasmid revealed potential SOS-mediated regulation of plasmid genes, highlighting the interaction between the SOS response and plasmid, although it did not significantly affect conjugation.IMPORTANCEPlasmids play a critical role in the dissemination of antibiotic resistance through conjugation. Recent research suggests that the use of antibiotics not only selects for already resistant variants but further increases the rate of plasmid-encoded conjugative transmission by increasing expression of the conjugative system. At the same time, these antibiotics are known to induce the stress-related SOS response in bacteria. To be able to counteract an antibiotic-induced increase in conjugative transfer of resistance plasmid, there is a need for a fundamental understanding of the regulation of transmission, including whether this happens through activation of the SOS response. In this research, we show that antibiotic-induced conjugation and induction of the SOS response happen through different mechanisms, and thus that future strategies to control the spread of antibiotics cannot interfere with the SOS response as its target.
Keywords: Escherichia coli; IncI1; SOS response; antibiotics; conjugation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Type I partition-related proteins enhance conjugative transfer through transcriptional activation and oriT region binding.mBio. 2025 Jul 31:e0160025. doi: 10.1128/mbio.01600-25. Online ahead of print. mBio. 2025. PMID: 40741764
-
Intestinal flora metabolites indole-3-butyric acid and disodium succinate promote IncI2 mcr-1-carrying plasmid transfer.Front Cell Infect Microbiol. 2025 Jun 3;15:1564810. doi: 10.3389/fcimb.2025.1564810. eCollection 2025. Front Cell Infect Microbiol. 2025. PMID: 40529306 Free PMC article.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2021 Apr 19;4(4):CD011535. doi: 10.1002/14651858.CD011535.pub4. Cochrane Database Syst Rev. 2021. Update in: Cochrane Database Syst Rev. 2022 May 23;5:CD011535. doi: 10.1002/14651858.CD011535.pub5. PMID: 33871055 Free PMC article. Updated.
-
Systemic pharmacological treatments for chronic plaque psoriasis: a network meta-analysis.Cochrane Database Syst Rev. 2017 Dec 22;12(12):CD011535. doi: 10.1002/14651858.CD011535.pub2. Cochrane Database Syst Rev. 2017. Update in: Cochrane Database Syst Rev. 2020 Jan 9;1:CD011535. doi: 10.1002/14651858.CD011535.pub3. PMID: 29271481 Free PMC article. Updated.
-
Interventions to improve antibiotic prescribing practices for hospital inpatients.Cochrane Database Syst Rev. 2017 Feb 9;2(2):CD003543. doi: 10.1002/14651858.CD003543.pub4. Cochrane Database Syst Rev. 2017. PMID: 28178770 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

