Gene Expression Analysis of (Paired) Primary and Relapsed Wilms Tumor Samples to Unravel the Underlying Factors Driving Tumor Recurrence
- PMID: 40439002
- PMCID: PMC12120524
- DOI: 10.1002/cam4.70969
Gene Expression Analysis of (Paired) Primary and Relapsed Wilms Tumor Samples to Unravel the Underlying Factors Driving Tumor Recurrence
Abstract
Purpose: We aimed to unravel underlying factors driving Wilms tumor (WT) recurrence and to build a prediction model for recurrence based on gene expression data of (paired) primary and relapsed WT samples.
Experimental design: Gene expression levels from seven paired primary and relapsed WT samples from patients treated in the Princess Máxima Center were compared among each other, as well as to matched primary WT samples of patients without recurrence (controls). The differential gene expression analysis results were run through ToppGene for functional enrichment. We built a 10-fold ridge regression model to predict relapse based on gene expression levels of the seven primary cases and all other available primary WT controls (n = 42).
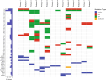
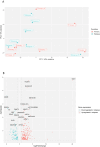
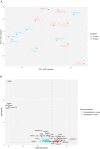
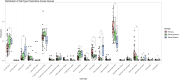
Results: The comparison of primary WT and paired relapses showed downregulation of genes involved in immune regulation among relapses and upregulation of cancer stem cell (CSC) regulation genes. Comparing these primary WT samples to matched controls, we observed that downregulated genes in primary samples of relapsed patients were related to stromal cells and muscle development, and upregulated genes were associated with CSCs. The prediction model revealed a sensitivity of 57.14% (95% CI: 14.29%-85.71%) and a specificity of 92.86% (95% CI: 83.33%-100%) when predicting WT relapse.
Conclusion: The CSC pool could play a role in relapse through immune regulation and tumor propagation. Differentiation of CSCs into mesenchymal cells might attenuate the risk of relapse. Our prediction model might aid in selecting patients with an increased risk of relapse at primary diagnosis when externally validated.
© 2025 The Author(s). Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
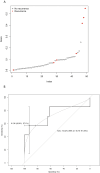
Figures

Similar articles
-
Construction of a Cancer Stem Cell Marker Genes-Related 22-Gene Signature for Overall Survival Prediction in High-Risk Wilms' Tumor.Fetal Pediatr Pathol. 2024 Sep-Oct;43(5):351-365. doi: 10.1080/15513815.2024.2382277. Epub 2024 Aug 6. Fetal Pediatr Pathol. 2024. PMID: 39108076
-
Chromosomal anomalies at 1q, 3, 16q, and mutations of SIX1 and DROSHA genes underlie Wilms tumor recurrences.Oncotarget. 2016 Feb 23;7(8):8908-15. doi: 10.18632/oncotarget.6950. Oncotarget. 2016. PMID: 26802027 Free PMC article.
-
Screening of potential hub genes involved in Kidney Wilms tumor via bioinformatics analysis and experimental validation.BMC Cancer. 2024 Jun 27;24(1):771. doi: 10.1186/s12885-024-12541-x. BMC Cancer. 2024. PMID: 38937666 Free PMC article.
-
Unmet needs for relapsed or refractory Wilms tumour: Mapping the molecular features, exploring organoids and designing early phase trials - A collaborative SIOP-RTSG, COG and ITCC session at the first SIOPE meeting.Eur J Cancer. 2021 Feb;144:113-122. doi: 10.1016/j.ejca.2020.11.012. Epub 2020 Dec 18. Eur J Cancer. 2021. PMID: 33341445 Review.
-
Wilms tumor--a renal stem cell malignancy?Pediatr Nephrol. 2011 Sep;26(9):1535-43. doi: 10.1007/s00467-011-1858-1. Epub 2011 Apr 16. Pediatr Nephrol. 2011. PMID: 21499773 Review.
References
-
- Brok J., Lopez‐Yurda M., Tinteren H. V., et al., “Relapse of Wilms' Tumour and Detection Methods: A Retrospective Analysis of the 2001 Renal Tumour Study Group‐International Society of Paediatric Oncology Wilms' Tumour Protocol Database,” Lancet Oncology 19, no. 8 (2018): 1072–1081, 10.1016/s1470-2045(18)30293-6. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

