Histone variant H2A.W7 represses meiotic crossover formation in Arabidopsis heterochromatin
- PMID: 40440068
- PMCID: PMC12146724
- DOI: 10.1073/pnas.2414166122
Histone variant H2A.W7 represses meiotic crossover formation in Arabidopsis heterochromatin
Abstract
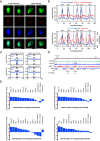
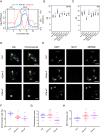
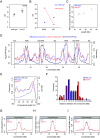
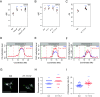
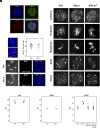
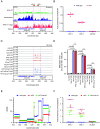
In eukaryotic genomes, DNA is packaged into nucleosomes to form chromatin. The incorporation of canonical or variant histones into nucleosomes confers different properties and influences chromatin structure to regulate cellular processes, including recombination. During meiosis, DNA double-strand breaks (DSBs) are formed and repaired as interhomolog crossovers. Nucleosome occupancy is generally associated with low crossover frequency, but it remains unclear which histone variants are involved in this process. In Arabidopsis, three variants of H2A coexist: H2A.X, H2A.Z, and H2A.W. Here, we show that H2A.W7 has a suppressive role on meiotic recombination. Genome-wide mapping of the crossover landscape revealed increased centromere-proximal recombination in h2a.w7. Moreover, H2A.W7 can be recruited to the 3a crossover hotspot via 21-24-nucleotide siRNAs during RNA-directed DNA methylation, causing increased nucleosome occupancy and decreased crossover frequency. Cytological analysis reveals that H2A.W7 restricts heterochromatin clustering during meiosis, which can form a mechanism to limit interhomolog recombination. Conversely, the linker histone H1, of which its loading is known to be restricted by H2A.W, promotes heterochromatin clustering and crossover on a heterochromatic genetic interval. Our study reveals a role for H2A.W7 in repressing crossover formation in Arabidopsis.
Keywords: H2A.W; crossover; epigenetics; meiosis; nucleosomes.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures
References
-
- Mercier R., Mézard C., Jenczewski E., Macaisne N., Grelon M., The molecular biology of meiosis in plants. Annu. Rev. Plant Biol. 66, 297–327 (2015). - PubMed
-
- Robert T., et al. , The TopoVIB-Like protein family is required for meiotic DNA double-strand break formation. Science. 351, 943–949 (2016). - PubMed
-
- Vrielynck N., et al. , A DNA topoisomerase VI-like complex initiates meiotic recombination. Science. 351, 939–943 (2016). - PubMed
-
- Bishop D. K., Park D., Xu L., Kleckner N., DMC1: A meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell. 69, 439–456 (1992). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

