Structural insights into light harvesting by antenna-containing rhodopsins in marine Asgard archaea
- PMID: 40442502
- PMCID: PMC12137139
- DOI: 10.1038/s41564-025-02016-5
Structural insights into light harvesting by antenna-containing rhodopsins in marine Asgard archaea
Abstract
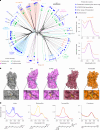
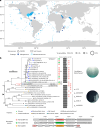
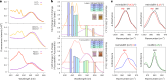
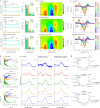
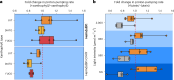
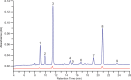
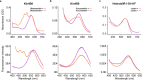
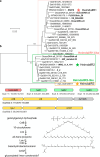
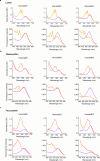
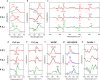
Aquatic bacterial rhodopsin proton pumps harvest light energy for photoheterotrophic growth and are known to contain hydroxylated carotenoids that expand the wavelengths of light utilized, but these have not been characterized in marine archaea. Here, by combining a marine chromophore extract with purified archaeal rhodopsins identified in marine metagenomes, we show light energy transfer from diverse hydroxylated carotenoids to heimdallarchaeial rhodopsins (HeimdallRs) from uncultured marine planktonic members of 'Candidatus Kariarchaeaceae' ('Candidatus Asgardarchaeota'). These light-harvesting antennas absorb in the blue-light range and transfer energy to the green-light-absorbing retinal chromophore within HeimdallRs, enabling the use of light that is otherwise unavailable to the rhodopsin. Furthermore, we show elevated proton pumping by the antennas in HeimdallRs under white-light illumination, which better simulates the light conditions encountered by these archaea in their natural habitats. Our results indicate that light-harvesting antennas in microbial rhodopsins exist in families beyond xanthorhodopsins and proteorhodopsins and are present in both marine bacteria and archaea.
© 2025. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures






Similar articles
-
Marine Bacterial and Archaeal Ion-Pumping Rhodopsins: Genetic Diversity, Physiology, and Ecology.Microbiol Mol Biol Rev. 2016 Sep 14;80(4):929-54. doi: 10.1128/MMBR.00003-16. Print 2016 Dec. Microbiol Mol Biol Rev. 2016. PMID: 27630250 Free PMC article. Review.
-
Phototrophy by antenna-containing rhodopsin pumps in aquatic environments.Nature. 2023 Mar;615(7952):535-540. doi: 10.1038/s41586-023-05774-6. Epub 2023 Mar 1. Nature. 2023. PMID: 36859551
-
Light-driven ion-translocating rhodopsins in marine bacteria.Trends Microbiol. 2015 Feb;23(2):91-8. doi: 10.1016/j.tim.2014.10.009. Trends Microbiol. 2015. PMID: 25432080 Review.
-
Xanthorhodopsin: Proton pump with a carotenoid antenna.Cell Mol Life Sci. 2007 Sep;64(18):2323-8. doi: 10.1007/s00018-007-7167-y. Cell Mol Life Sci. 2007. PMID: 17571211 Free PMC article. Review.
-
Xanthorhodopsin: a proton pump with a light-harvesting carotenoid antenna.Science. 2005 Sep 23;309(5743):2061-4. doi: 10.1126/science.1118046. Science. 2005. PMID: 16179480 Free PMC article.
References
-
- Rozenberg, A., Inoue, K., Kandori, H. & Béjà, O. Microbial rhodopsins: the last two decades. Annu. Rev. Microbiol.75, 427–447 (2021). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

